Prostate-derived circulating microRNAs add prognostic value to prostate cancer risk calculators
- PMID: 38496750
- PMCID: PMC10938556
- DOI: 10.1002/jex2.122
Prostate-derived circulating microRNAs add prognostic value to prostate cancer risk calculators
Abstract
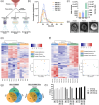
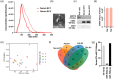
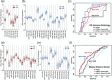
Prostate cancer is the second leading cause of malignancy-related deaths among American men. Active surveillance is a safe option for many men with less aggressive disease, yet definitively determining low-risk cancer is challenging with biopsy alone. Herein, we sought to identify prostate-derived microRNAs in patient sera and serum extracellular vesicles, and determine if those microRNAs improve upon the current clinical risk calculators for prostate cancer prognosis before and after biopsy. Prostate-derived intracellular and extracellular vesicle-contained microRNAs were identified by small RNA sequencing of prostate cancer patient explants and primary cells. Abundant microRNAs were included in a custom microRNA PCR panel that was queried in whole serum and serum extracellular vesicles from a diverse cohort of men diagnosed with prostate cancer. The levels of these circulating microRNAs significantly differed between indolent and aggressive disease and improved the area under the curve for pretreatment nomograms of prostate cancer disease risk. The microRNAs within the extracellular vesicles were the most informative and improved the AUC to 0.739 compared to the existing nomogram alone, which has an AUC of 0.561. The microRNAs in the whole serum improved it to AUC 0.675. In summary, quantifying microRNAs circulating in extracellular vesicles is a clinically feasible assay that may provide additional information for assessing prostate cancer risk stratification.
Conflict of interest statement
CONFLICT OF INTEREST DECLARATION The authors declare that they have no affiliations with or involvement in any organization or entity with any financial interest in the subject matter or materials discussed in this manuscript. The authors have declared that no conflict of interest exists.
Figures

Update of
-
Prostate-derived circulating microRNAs add prognostic value to prostate cancer risk calculators.bioRxiv [Preprint]. 2023 Jun 6:2023.05.10.540236. doi: 10.1101/2023.05.10.540236. bioRxiv. 2023. Update in: J Extracell Biol. 2023 Nov;2(11):e122. doi: 10.1002/jex2.122. PMID: 37214878 Free PMC article. Updated. Preprint.
References
-
- Aalberts, M. , Stout, T. A. , & Stoorvogel, W. (2014). Prostasomes: Extracellular vesicles from the prostate. Reproduction (Cambridge, England), 147, R1–14. - PubMed
-
- Aas, K. , Dorothea Fosså, S. , Åge Myklebust, T. , Møller, B. , Kvåle, R. , Vlatkovic, L. , & Berge, V. (2020). Increased curative treatment is associated with decreased prostate cancer‐specific and overall mortality in senior adults with high‐risk prostate cancer; results from a national registry‐based cohort study. Cancer Medicine, 9(18), 6646–6657. 10.1002/cam4.3297 - DOI - PMC - PubMed
-
- Ankerst, D. P. , Straubinger, J. , Selig, K. , Guerrios, L. , De Hoedt, A. , Hernandez, J. , Liss, M. A. , Leach, R. J. , Freedland, S. J. , Kattan, M. W. , Nam, R. , Haese, A. , Montorsi, F. , Boorjian, S. A. , Cooperberg, M. R. , Poyet, C. , Vertosick, E. , & Vickers, A. J. (2018). A contemporary prostate biopsy risk calculator based on multiple heterogeneous cohorts. European Urology, 74(2), 197–203. 10.1016/j.eururo.2018.05.003 - DOI - PMC - PubMed
-
- Benjamini, Y. , & Hochberg, Y. (1995). Controlling the false discovery rate: A practical and powerful approach to multiple testing. Journal of the Royal Statistical Society: Series B (Methodological), 57, 289–300.
Grants and funding
LinkOut - more resources
Full Text Sources
