Generation of marmoset primordial germ cell-like cells under chemically defined conditions
- PMID: 38499329
- PMCID: PMC10948935
- DOI: 10.26508/lsa.202302371
Generation of marmoset primordial germ cell-like cells under chemically defined conditions
Abstract
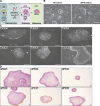
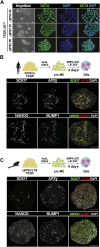
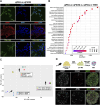
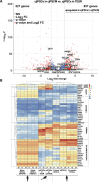
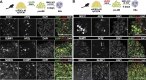
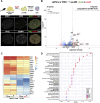
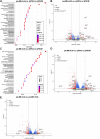
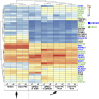
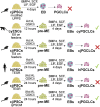
Primordial germ cells (PGCs) are the embryonic precursors of sperm and oocytes, which transmit genetic/epigenetic information across generations. Mouse PGC and subsequent gamete development can be fully reconstituted in vitro, opening up new avenues for germ cell studies in biomedical research. However, PGCs show molecular differences between rodents and humans. Therefore, to establish an in vitro system that is closely related to humans, we studied PGC development in vivo and in vitro in the common marmoset monkey Callithrix jacchus (cj). Gonadal cjPGCs at embryonic day 74 express SOX17, AP2Ɣ, BLIMP1, NANOG, and OCT4A, which is reminiscent of human PGCs. We established transgene-free induced pluripotent stem cell (cjiPSC) lines from foetal and postnatal fibroblasts. These cjiPSCs, cultured in defined and feeder-free conditions, can be differentiated into precursors of mesendoderm and subsequently into cjPGC-like cells (cjPGCLCs) with a transcriptome similar to human PGCs/PGCLCs. Our results not only pave the way for studying PGC development in a non-human primate in vitro under experimentally controlled conditions, but also provide the opportunity to derive functional marmoset gametes in future studies.
© 2024 Kurlovich et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures






Similar articles
-
Efficient generation of marmoset primordial germ cell-like cells using induced pluripotent stem cells.Elife. 2023 Jan 31;12:e82263. doi: 10.7554/eLife.82263. Elife. 2023. PMID: 36719274 Free PMC article.
-
Primordial germ cell development in the marmoset monkey as revealed by pluripotency factor expression: suggestion of a novel model of embryonic germ cell translocation.Mol Hum Reprod. 2015 Jan;21(1):66-80. doi: 10.1093/molehr/gau088. Epub 2014 Sep 18. Mol Hum Reprod. 2015. PMID: 25237007 Free PMC article.
-
Making gametes from pluripotent stem cells--a promising role for very small embryonic-like stem cells.Reprod Biol Endocrinol. 2014 Nov 24;12:114. doi: 10.1186/1477-7827-12-114. Reprod Biol Endocrinol. 2014. PMID: 25421462 Free PMC article.
-
How to make a primordial germ cell.Development. 2014 Jan;141(2):245-52. doi: 10.1242/dev.098269. Development. 2014. PMID: 24381195 Review.
-
From stem cells to spermatozoa and back.Soc Reprod Fertil Suppl. 2007;65:19-32. Soc Reprod Fertil Suppl. 2007. PMID: 17644952 Review.
Cited by
-
Stem cell-based embryo models: a tool to study early human development.Sci China Life Sci. 2025 Jun;68(6):1626-1645. doi: 10.1007/s11427-024-2741-1. Epub 2025 Feb 17. Sci China Life Sci. 2025. PMID: 39969747 Review.
References
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
