Cryo-EM structures of type IV pili complexed with nanobodies reveal immune escape mechanisms
- PMID: 38499587
- PMCID: PMC10948894
- DOI: 10.1038/s41467-024-46677-y
Cryo-EM structures of type IV pili complexed with nanobodies reveal immune escape mechanisms
Erratum in
-
Publisher Correction: Cryo-EM structures of type IV pili complexed with nanobodies reveal immune escape mechanisms.Nat Commun. 2024 Apr 3;15(1):2873. doi: 10.1038/s41467-024-47081-2. Nat Commun. 2024. PMID: 38570508 Free PMC article. No abstract available.
Abstract
Type IV pili (T4P) are prevalent, polymeric surface structures in pathogenic bacteria, making them ideal targets for effective vaccines. However, bacteria have evolved efficient strategies to evade type IV pili-directed antibody responses. Neisseria meningitidis are prototypical type IV pili-expressing Gram-negative bacteria responsible for life threatening sepsis and meningitis. This species has evolved several genetic strategies to modify the surface of its type IV pili, changing pilin subunit amino acid sequence, nature of glycosylation and phosphoforms, but how these modifications affect antibody binding at the structural level is still unknown. Here, to explore this question, we determine cryo-electron microscopy (cryo-EM) structures of pili of different sequence types with sufficiently high resolution to visualize posttranslational modifications. We then generate nanobodies directed against type IV pili which alter pilus function in vitro and in vivo. Cyro-EM in combination with molecular dynamics simulation of the nanobody-pilus complexes reveals how the different types of pili surface modifications alter nanobody binding. Our findings shed light on the impressive complementarity between the different strategies used by bacteria to avoid antibody binding. Importantly, we also show that structural information can be used to make informed modifications in nanobodies as countermeasures to these immune evasion mechanisms.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interest.
Figures


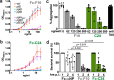




References
-
- Ellison, C. K., Whitfield, G. B. & Brun, Y. V. Type IV Pili: Dynamic bacterial nanomachines. FEMS Microbiol. Rev. fuab053 10.1093/femsre/fuab053 (2021). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

