Towards a general-purpose foundation model for computational pathology
- PMID: 38504018
- PMCID: PMC11403354
- DOI: 10.1038/s41591-024-02857-3
Towards a general-purpose foundation model for computational pathology
Abstract
Quantitative evaluation of tissue images is crucial for computational pathology (CPath) tasks, requiring the objective characterization of histopathological entities from whole-slide images (WSIs). The high resolution of WSIs and the variability of morphological features present significant challenges, complicating the large-scale annotation of data for high-performance applications. To address this challenge, current efforts have proposed the use of pretrained image encoders through transfer learning from natural image datasets or self-supervised learning on publicly available histopathology datasets, but have not been extensively developed and evaluated across diverse tissue types at scale. We introduce UNI, a general-purpose self-supervised model for pathology, pretrained using more than 100 million images from over 100,000 diagnostic H&E-stained WSIs (>77 TB of data) across 20 major tissue types. The model was evaluated on 34 representative CPath tasks of varying diagnostic difficulty. In addition to outperforming previous state-of-the-art models, we demonstrate new modeling capabilities in CPath such as resolution-agnostic tissue classification, slide classification using few-shot class prototypes, and disease subtyping generalization in classifying up to 108 cancer types in the OncoTree classification system. UNI advances unsupervised representation learning at scale in CPath in terms of both pretraining data and downstream evaluation, enabling data-efficient artificial intelligence models that can generalize and transfer to a wide range of diagnostically challenging tasks and clinical workflows in anatomic pathology.
© 2024. The Author(s), under exclusive licence to Springer Nature America, Inc.
Figures




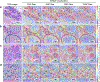
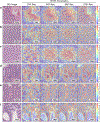


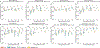


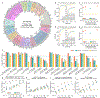


References
-
- Song AH et al. Artificial intelligence for digital and computational pathology. Nat. Rev. Bioeng. 1, 930–949 (2023).
-
- Heinz CN, Echle A, Foersch S, Bychkov A & Kather JN The future of artificial intelligence in digital pathology: results of a survey across stakeholder groups. Histopathology 80, 1121–1127 (2022). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

