PROTAC-mediated degradation of HIV-1 Nef efficiently restores cell-surface CD4 and MHC-I expression and blocks HIV-1 replication
- PMID: 38508197
- PMCID: PMC11031313
- DOI: 10.1016/j.chembiol.2024.02.004
PROTAC-mediated degradation of HIV-1 Nef efficiently restores cell-surface CD4 and MHC-I expression and blocks HIV-1 replication
Abstract
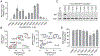
The HIV-1 Nef accessory factor enhances the viral life cycle in vivo, promotes immune escape of HIV-infected cells, and represents an attractive antiretroviral drug target. However, Nef lacks enzymatic activity and an active site, complicating traditional occupancy-based drug development. Here we describe the development of proteolysis targeting chimeras (PROTACs) for the targeted degradation of Nef. Nef-binding compounds, based on an existing hydroxypyrazole core, were coupled to ligands for ubiquitin E3 ligases via flexible linkers. The resulting bivalent PROTACs induced formation of a ternary complex between Nef and the cereblon E3 ubiquitin ligase thalidomide-binding domain in vitro and triggered Nef degradation in a T cell expression system. Nef-directed PROTACs efficiently rescued Nef-mediated MHC-I and CD4 downregulation in T cells and suppressed HIV-1 replication in donor PBMCs. Targeted degradation is anticipated to reverse all HIV-1 Nef functions and may help restore adaptive immune responses against HIV-1 reservoir cells in vivo.
Keywords: CD4; HIV Nef; HIV-1; MHC-I; PROTACs; antiretroviral drugs; targeted degradation.
Copyright © 2024 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures



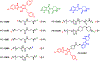


Update of
-
PROTAC-mediated Degradation of HIV-1 Nef Efficiently Restores Cell-surface CD4 and MHC-I Expression and Blocks HIV-1 Replication.bioRxiv [Preprint]. 2023 Sep 5:2023.08.14.553289. doi: 10.1101/2023.08.14.553289. bioRxiv. 2023. Update in: Cell Chem Biol. 2024 Apr 18;31(4):658-668.e14. doi: 10.1016/j.chembiol.2024.02.004. PMID: 37645900 Free PMC article. Updated. Preprint.
Comment in
-
PROTAC induces HIV-1 Nef degradation.Nat Rev Drug Discov. 2024 May;23(5):340. doi: 10.1038/d41573-024-00060-0. Nat Rev Drug Discov. 2024. PMID: 38575781 No abstract available.
References
-
- Bregnard C, Zamborlini A, Leduc M, Chafey P, Camoin L, Saib A, Benichou S, Danos O, and Basmaciogullari S (2013). Comparative proteomic analysis of HIV-1 particles reveals a role for Ezrin and EHD4 in the Nef-dependent increase of virus infectivity. J. Virol 87, 3729–3740. JVI.02477-12 [pii]; 10.1128/JVI.02477-12 [doi]. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

