Disease-associated astrocyte epigenetic memory promotes CNS pathology
- PMID: 38509377
- PMCID: PMC11016191
- DOI: 10.1038/s41586-024-07187-5
Disease-associated astrocyte epigenetic memory promotes CNS pathology
Erratum in
-
Author Correction: Disease-associated astrocyte epigenetic memory promotes CNS pathology.Nature. 2024 Jul;631(8020):E8. doi: 10.1038/s41586-024-07734-0. Nature. 2024. PMID: 38907058 No abstract available.
-
Author Correction: Disease-associated astrocyte epigenetic memory promotes CNS pathology.Nature. 2025 Sep;645(8081):E5. doi: 10.1038/s41586-025-09546-2. Nature. 2025. PMID: 40858938 No abstract available.
Abstract
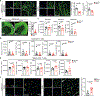
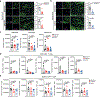
Disease-associated astrocyte subsets contribute to the pathology of neurologic diseases, including multiple sclerosis and experimental autoimmune encephalomyelitis1-8 (EAE), an experimental model for multiple sclerosis. However, little is known about the stability of these astrocyte subsets and their ability to integrate past stimulation events. Here we report the identification of an epigenetically controlled memory astrocyte subset that exhibits exacerbated pro-inflammatory responses upon rechallenge. Specifically, using a combination of single-cell RNA sequencing, assay for transposase-accessible chromatin with sequencing, chromatin immunoprecipitation with sequencing, focused interrogation of cells by nucleic acid detection and sequencing, and cell-specific in vivo CRISPR-Cas9-based genetic perturbation studies we established that astrocyte memory is controlled by the metabolic enzyme ATP-citrate lyase (ACLY), which produces acetyl coenzyme A (acetyl-CoA) that is used by histone acetyltransferase p300 to control chromatin accessibility. The number of ACLY+p300+ memory astrocytes is increased in acute and chronic EAE models, and their genetic inactivation ameliorated EAE. We also detected the pro-inflammatory memory phenotype in human astrocytes in vitro; single-cell RNA sequencing and immunohistochemistry studies detected increased numbers of ACLY+p300+ astrocytes in chronic multiple sclerosis lesions. In summary, these studies define an epigenetically controlled memory astrocyte subset that promotes CNS pathology in EAE and, potentially, multiple sclerosis. These findings may guide novel therapeutic approaches for multiple sclerosis and other neurologic diseases.
© 2024. The Author(s), under exclusive licence to Springer Nature Limited.
Conflict of interest statement
Figures













Update of
-
Disease-associated astrocyte epigenetic memory promotes CNS pathology.bioRxiv [Preprint]. 2024 Jan 5:2024.01.04.574196. doi: 10.1101/2024.01.04.574196. bioRxiv. 2024. Update in: Nature. 2024 Mar;627(8005):865-872. doi: 10.1038/s41586-024-07187-5. PMID: 38260616 Free PMC article. Updated. Preprint.
Comment in
-
Astrocytes remember inflammation.Immunity. 2024 May 14;57(5):938-940. doi: 10.1016/j.immuni.2024.04.005. Immunity. 2024. PMID: 38749396
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

