Dynamic immune recovery process after liver transplantation revealed by single-cell multi-omics analysis
- PMID: 38510071
- PMCID: PMC10952083
- DOI: 10.1016/j.xinn.2024.100599
Dynamic immune recovery process after liver transplantation revealed by single-cell multi-omics analysis
Abstract
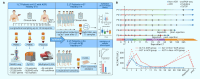
Elucidating the temporal process of immune remodeling under immunosuppressive treatment after liver transplantation (LT) is critical for precise clinical management strategies. Here, we performed a single-cell multi-omics analysis of peripheral blood mononuclear cells (PBMCs) collected from LT patients (with and without acute cellular rejection [ACR]) at 13 time points. Validation was performed in two independent cohorts with additional LT patients and healthy controls. Our study revealed a four-phase recovery process after LT and delineated changes in immune cell composition, expression programs, and interactions along this process. The intensity of the immune response differs between the ACR and non-ACR patients. Notably, the newly identified inflamed NK cells, CD14+RNASE2+ monocytes, and FOS-expressing monocytes emerged as predictive indicators of ACR. This study illuminates the longitudinal evolution of the immune cell landscape under tacrolimus-based immunosuppressive treatment during LT recovery, providing a four-phase framework that aids the clinical management of LT patients.
© 2024 The Authors.
Conflict of interest statement
The authors declare no competing interests.
Figures








References
LinkOut - more resources
Full Text Sources
Research Materials

