Type I and II interferons, transcription factors and major histocompatibility complexes were enhanced by knocking down the PRRSV-induced transforming growth factor beta in monocytes co-cultured with peripheral blood lymphocytes
- PMID: 38510257
- PMCID: PMC10950996
- DOI: 10.3389/fimmu.2024.1308330
Type I and II interferons, transcription factors and major histocompatibility complexes were enhanced by knocking down the PRRSV-induced transforming growth factor beta in monocytes co-cultured with peripheral blood lymphocytes
Abstract
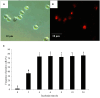
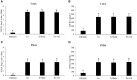
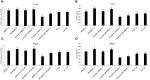
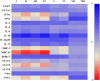
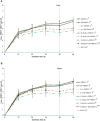
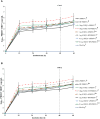
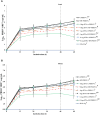
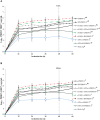
The innate and adaptive immune responses elicited by porcine reproductive and respiratory syndrome virus (PRRSV) infection are known to be poor. This study investigates the impact of PRRSV-induced transforming growth factor beta 1 (TGFβ1) on the expressions of type I and II interferons (IFNs), transcription factors, major histocompatibility complexes (MHC), anti-inflammatory and pro-inflammatory cytokines in PRRSV-infected co-cultures of monocytes and peripheral blood lymphocytes (PBL). Phosphorothioate-modified antisense oligodeoxynucleotide (AS ODN) specific to the AUG region of porcine TGFβ1 mRNA was synthesized and successfully knocked down TGFβ1 mRNA expression and protein translation. Monocytes transfected with TGFβAS1 ODN, then simultaneously co-cultured with PBL and inoculated with either classical PRRSV-2 (cPRRSV-2) or highly pathogenic PRRSV-2 (HP-PRRSV-2) showed a significant reduction in TGFβ1 mRNA expression and a significant increase in the mRNA expressions of IFNα, IFNγ, MHC-I, MHC-II, signal transducer and activator of transcription 1 (STAT1), and STAT2. Additionally, transfection of TGFβAS1 ODN in the monocyte and PBL co-culture inoculated with cPRRSV-2 significantly increased the mRNA expression of interleukin-12p40 (IL-12p40). PRRSV-2 RNA copy numbers were significantly reduced in monocytes and PBL co-culture transfected with TGFβAS1 ODN compared to the untransfected control. The yields of PRRSV-2 RNA copy numbers in PRRSV-2-inoculated monocytes and PBL co-culture were sustained and reduced by porcine TGFβ1 (rTGFβ1) and recombinant porcine IFNα (rIFNα), respectively. These findings highlight the strategy employed by PRRSV to suppress the innate immune response through the induction of TGFβ expression. The inclusion of TGFβ as a parameter for future PRRSV vaccine and vaccine adjuvant candidates is recommended.
Keywords: antisense; innate immunity; porcine reproductive and respiratory syndrome virus; transforming growth factor beta; type I and II interferons.
Copyright © 2024 Fabros and Charerntantanakul.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Knock down of transforming growth factor beta improves expressions of co-stimulatory molecules, type I interferon-regulated genes, and pro-inflammatory cytokine in PRRSV-inoculated monocyte-derived macrophages.BMC Vet Res. 2024 Aug 3;20(1):344. doi: 10.1186/s12917-023-03760-8. BMC Vet Res. 2024. PMID: 39097704 Free PMC article.
-
Plasmids expressing porcine interferon gamma up-regulate pro-inflammatory cytokine and co-stimulatory molecule expression which are suppressed by porcine reproductive and respiratory syndrome virus.Vet Immunol Immunopathol. 2013 May 15;153(1-2):107-17. doi: 10.1016/j.vetimm.2013.02.013. Epub 2013 Feb 26. Vet Immunol Immunopathol. 2013. PMID: 23507439
-
Rutin, α-tocopherol, and l-ascorbic acid up-regulate type I interferon-regulated gene and type I and II interferon expressions and reduce inflammatory cytokine expressions in monocyte-derived macrophages infected with highly pathogenic porcine reproductive and respiratory syndrome virus.Vet Immunol Immunopathol. 2021 May;235:110231. doi: 10.1016/j.vetimm.2021.110231. Epub 2021 Mar 15. Vet Immunol Immunopathol. 2021. PMID: 33740613
-
Saponin Quil A up-regulates type I interferon-regulated gene and type I and II interferon expressions which are suppressed by porcine reproductive and respiratory syndrome virus.Vet Immunol Immunopathol. 2018 Jan;195:76-83. doi: 10.1016/j.vetimm.2017.11.009. Epub 2017 Dec 6. Vet Immunol Immunopathol. 2018. PMID: 29249322
-
Adjuvants for porcine reproductive and respiratory syndrome virus vaccines.Vet Immunol Immunopathol. 2009 May 15;129(1-2):1-13. doi: 10.1016/j.vetimm.2008.12.018. Epub 2008 Dec 11. Vet Immunol Immunopathol. 2009. PMID: 19157569 Review.
References
-
- Zimmerman JJ, Dee SA, Holtkamp DJ, Murtaugh MP, Stadejek T, Stevenson GW, et al. . Porcine reproductive and respiratory syndrome viruses (porcine arteriviruses). Dis swine. (2019), 685–708. doi: 10.1002/9781119350927.ch41 - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

