A genome-wide CRISPR screening uncovers that TOB1 acts as a key host factor for FMDV infection via both IFN and EGFR mediated pathways
- PMID: 38512977
- PMCID: PMC10986976
- DOI: 10.1371/journal.ppat.1012104
A genome-wide CRISPR screening uncovers that TOB1 acts as a key host factor for FMDV infection via both IFN and EGFR mediated pathways
Abstract
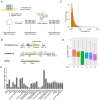
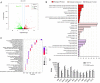
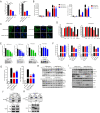
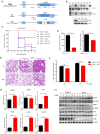
The interaction between foot-and-mouth disease virus (FMDV) and the host is extremely important for virus infection, but there are few researches on it, which is not conducive to vaccine development and FMD control. In this study, we designed a porcine genome-scale CRISPR/Cas9 knockout library containing 93,859 single guide RNAs targeting 16,886 protein-coding genes, 25 long ncRNAs, and 463 microRNAs. Using this library, several previously unreported genes required for FMDV infection are highly enriched post-FMDV selection in IBRS-2 cells. Follow-up studies confirmed the dependency of FMDV on these genes, and we identified a functional role for one of the FMDV-related host genes: TOB1 (Transducer of ERBB2.1). TOB1-knockout significantly inhibits FMDV infection by positively regulating the expression of RIG-I and MDA5. We further found that TOB1-knockout led to more accumulation of mRNA transcripts of transcription factor CEBPA, and thus its protein, which further enhanced transcription of RIG-I and MDA5 genes. In addition, TOB1-knockout was shown to inhibit FMDV adsorption and internalization mediated by EGFR/ERBB2 pathway. Finally, the FMDV lethal challenge on TOB1-knockout mice confirmed that the deletion of TOB1 inhibited FMDV infection in vivo. These results identify TOB1 as a key host factor involved in FMDV infection in pigs.
Copyright: © 2024 Peng et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

