Activation of ERβ hijacks the splicing machinery to trigger R-loop formation in triple-negative breast cancer
- PMID: 38513102
- PMCID: PMC10990146
- DOI: 10.1073/pnas.2306814121
Activation of ERβ hijacks the splicing machinery to trigger R-loop formation in triple-negative breast cancer
Abstract
Triple-negative breast cancer (TNBC) is a subtype of breast cancer with aggressive behavior and poor prognosis. Current therapeutic options available for TNBC patients are primarily chemotherapy. With our evolving understanding of this disease, novel targeted therapies, including poly ADP-ribose polymerase (PARP) inhibitors, antibody-drug conjugates, and immune-checkpoint inhibitors, have been developed for clinical use. Previous reports have demonstrated the essential role of estrogen receptor β (ERβ) in TNBC, but the detailed molecular mechanisms downstream ERβ activation in TNBC are still far from elucidated. In this study, we demonstrated that a specific ERβ agonist, LY500307, potently induces R-loop formation and DNA damage in TNBC cells. Subsequent interactome experiments indicated that the residues 151 to 165 of U2 small nuclear RNA auxiliary factor 1 (U2AF1) and the Trp439 and Lys443 of ERβ were critical for the binding between U2AF1 and ERβ. Combined RNA sequencing and ribosome sequencing analysis demonstrated that U2AF1-regulated downstream RNA splicing of 5-oxoprolinase (OPLAH) could affect its enzymatic activity and is essential for ERβ-induced R-loop formation and DNA damage. In clinical samples including 115 patients from The Cancer Genome Atlas (TCGA) and 32 patients from an in-house cohort, we found a close correlation in the expression of ESR2 and U2AF1 in TNBC patients. Collectively, our study has unraveled the molecular mechanisms that explain the therapeutic effects of ERβ activation in TNBC, which provides rationale for ERβ activation-based single or combined therapy for patients with TNBC.
Keywords: ERβ; nuclear receptors; triple-negative breast cancer.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures
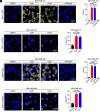
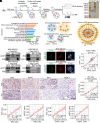
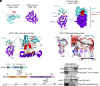

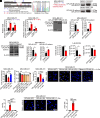
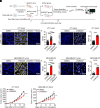
Similar articles
-
Pharmacological activation of estrogen receptor beta augments innate immunity to suppress cancer metastasis.Proc Natl Acad Sci U S A. 2018 Apr 17;115(16):E3673-E3681. doi: 10.1073/pnas.1803291115. Epub 2018 Mar 28. Proc Natl Acad Sci U S A. 2018. PMID: 29592953 Free PMC article.
-
Epigenetic restoration and activation of ERβ: an inspiring approach for treatment of triple-negative breast cancer.Med Oncol. 2022 Jul 18;39(10):150. doi: 10.1007/s12032-022-01765-1. Med Oncol. 2022. PMID: 35843988 Free PMC article.
-
Interaction Proteomics Identifies ERbeta Association with Chromatin Repressive Complexes to Inhibit Cholesterol Biosynthesis and Exert An Oncosuppressive Role in Triple-negative Breast Cancer.Mol Cell Proteomics. 2020 Feb;19(2):245-260. doi: 10.1074/mcp.RA119.001817. Epub 2019 Dec 2. Mol Cell Proteomics. 2020. PMID: 31792072 Free PMC article.
-
Biology and Management of Patients With Triple-Negative Breast Cancer.Oncologist. 2016 Sep;21(9):1050-62. doi: 10.1634/theoncologist.2016-0067. Epub 2016 Jul 11. Oncologist. 2016. PMID: 27401886 Free PMC article. Review.
-
Perspectives on Triple-Negative Breast Cancer: Current Treatment Strategies, Unmet Needs, and Potential Targets for Future Therapies.Cancers (Basel). 2020 Aug 24;12(9):2392. doi: 10.3390/cancers12092392. Cancers (Basel). 2020. PMID: 32846967 Free PMC article. Review.
Cited by
-
Effects of Garlic on Breast Tumor Cells with a Triple Negative Phenotype: Peculiar Subtype-Dependent Down-Modulation of Akt Signaling.Cells. 2024 May 11;13(10):822. doi: 10.3390/cells13100822. Cells. 2024. PMID: 38786044 Free PMC article.
-
Mechanisms of RNA alternative splicing dysregulation in triple-negative breast cancer.Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2024 Jul 28;49(7):1143-1154. doi: 10.11817/j.issn.1672-7347.2024.240434. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2024. PMID: 39788502 Free PMC article. Review. Chinese, English.
-
Androgen receptor expression and clinical characteristics in breast cancer.World J Surg Oncol. 2024 Sep 11;22(1):243. doi: 10.1186/s12957-024-03525-z. World J Surg Oncol. 2024. PMID: 39256855 Free PMC article.
-
Identifying PSIP1 as a critical R-loop regulator in osteosarcoma via machine-learning and multi-omics analysis.Cancer Cell Int. 2025 Apr 22;25(1):159. doi: 10.1186/s12935-025-03775-1. Cancer Cell Int. 2025. PMID: 40264164 Free PMC article.
References
-
- Bianchini G., De Angelis C., Licata L., Gianni L., Treatment landscape of triple-negative breast cancer–Expanded options, evolving needs. Nat. Rev. Clin. Oncol. 19, 91–113 (2022). - PubMed
-
- Hartman J., et al. , Estrogen receptor beta inhibits angiogenesis and growth of T47D breast cancer xenografts. Cancer Res. 66, 11207–11213 (2006). - PubMed
-
- Honma N., et al. , Clinical importance of estrogen receptor-beta evaluation in breast cancer patients treated with adjuvant tamoxifen therapy. J. Clin. Oncol. 26, 3727–3734 (2008). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

