The histone acetyltransferase KAT6B is required for hematopoietic stem cell development and function
- PMID: 38518784
- PMCID: PMC11096436
- DOI: 10.1016/j.stemcr.2024.02.005
The histone acetyltransferase KAT6B is required for hematopoietic stem cell development and function
Abstract
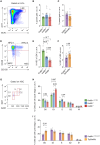
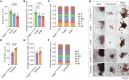
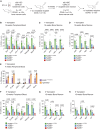
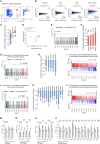
The histone lysine acetyltransferase KAT6B (MYST4, MORF, QKF) is the target of recurrent chromosomal translocations causing hematological malignancies with poor prognosis. Using Kat6b germline deletion and overexpression in mice, we determined the role of KAT6B in the hematopoietic system. We found that KAT6B sustained the fetal hematopoietic stem cell pool but did not affect viability or differentiation. KAT6B was essential for normal levels of histone H3 lysine 9 (H3K9) acetylation but not for a previously proposed target, H3K23. Compound heterozygosity of Kat6b and the closely related gene, Kat6a, abolished hematopoietic reconstitution after transplantation. KAT6B and KAT6A cooperatively promoted transcription of genes regulating hematopoiesis, including the Hoxa cluster, Pbx1, Meis1, Gata family, Erg, and Flt3. In conclusion, we identified the hematopoietic processes requiring Kat6b and showed that KAT6B and KAT6A synergistically promoted HSC development, function, and transcription. Our findings are pertinent to current clinical trials testing KAT6A/B inhibitors as cancer therapeutics.
Keywords: KAT6A; KAT6B; chromatin; hematopoiesis; histone acetyltransferase; stem cells; transplantation.
Copyright © 2024 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests T.T. and A.K.V. are inventors on patent WO2016198507 A1. The Thomas and Voss laboratories received research funding from the CRC for Cancer Therapeutics (CTx), Australia. T.T. and A.K.V. have received payments from a distribution of licensing income from Pfizer and have served on an advisory board for Pfizer.
Figures



References
-
- Adelman E.R., Huang H.T., Roisman A., Olsson A., Colaprico A., Qin T., Lindsley R.C., Bejar R., Salomonis N., Grimes H.L., Figueroa M.E. Aging human hematopoietic stem cells manifest profound epigenetic reprogramming of enhancers that may predispose to leukemia. Cancer Discov. 2019;9:1080–1101. - PMC - PubMed
-
- Arai F., Hirao A., Ohmura M., Sato H., Matsuoka S., Takubo K., Ito K., Koh G.Y., Suda T. Tie2/angiopoietin-1 signaling regulates hematopoietic stem cell quiescence in the bone marrow niche. Cell. 2004;118:149–161. - PubMed
-
- Baell J.B., Leaver D.J., Hermans S.J., Kelly G.L., Brennan M.S., Downer N.L., Nguyen N., Wichmann J., McRae H.M., Yang Y., et al. Inhibitors of histone acetyltransferases KAT6A/B induce senescence and arrest tumour growth. Nature. 2018;560:253–257. - PubMed
-
- Borrow J., Stanton V.P., Jr., Andresen J.M., Becher R., Behm F.G., Chaganti R.S., Civin C.I., Disteche C., Dubé I., Frischauf A.M., et al. The translocation t(8;16)(p11;p13) of acute myeloid leukaemia fuses a putative acetyltransferase to the CREB-binding protein. Nat. Genet. 1996;14:33–41. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

