Genetic contribution to microglial activation in schizophrenia
- PMID: 38519640
- PMCID: PMC11420079
- DOI: 10.1038/s41380-024-02529-1
Genetic contribution to microglial activation in schizophrenia
Abstract
Several lines of evidence indicate the involvement of neuroinflammatory processes in the pathophysiology of schizophrenia (SCZ). Microglia are brain resident immune cells responding toward invading pathogens and injury-related products, and additionally, have a critical role in improving neurogenesis and synaptic functions. Aberrant activation of microglia in SCZ is one of the leading hypotheses for disease pathogenesis, but due to the lack of proper human cell models, the role of microglia in SCZ is not well studied. We used monozygotic twins discordant for SCZ and healthy individuals to generate human induced pluripotent stem cell-derived microglia to assess the transcriptional and functional differences in microglia between healthy controls, affected twins and unaffected twins. The microglia from affected twins had increased expression of several common inflammation-related genes compared to healthy individuals. Microglia from affected twins had also reduced response to interleukin 1 beta (IL1β) treatment, but no significant differences in migration or phagocytotic activity. Ingenuity Pathway Analysis (IPA) showed abnormalities related to extracellular matrix signaling. RNA sequencing predicted downregulation of extracellular matrix structure constituent Gene Ontology (GO) terms and hepatic fibrosis pathway activation that were shared by microglia of both affected and unaffected twins, but the upregulation of major histocompatibility complex (MHC) class II receptors was observed only in affected twin microglia. Also, the microglia of affected twins had heterogeneous response to clozapine, minocycline, and sulforaphane treatments. Overall, despite the increased expression of inflammatory genes, we observed no clear functional signs of hyperactivation in microglia from patients with SCZ. We conclude that microglia of the patients with SCZ have gene expression aberrations related to inflammation response and extracellular matrix without contributing to increased microglial activation.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





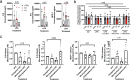
References
-
- Monji A, Kato T, Kanba S. Cytokines and schizophrenia: microglia hypothesis of schizophrenia: cytokines and schizophrenia. Psychiatry Clin Neurosci. 2009;63:257–65. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

