Diverse functions of cytochrome c in cell death and disease
- PMID: 38521844
- PMCID: PMC11043370
- DOI: 10.1038/s41418-024-01284-8
Diverse functions of cytochrome c in cell death and disease
Abstract
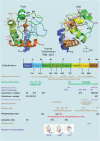
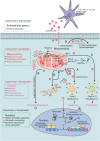
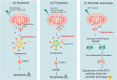
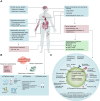
The redox-active protein cytochrome c is a highly positively charged hemoglobin that regulates cell fate decisions of life and death. Under normal physiological conditions, cytochrome c is localized in the mitochondrial intermembrane space, and its distribution can extend to the cytosol, nucleus, and extracellular space under specific pathological or stress-induced conditions. In the mitochondria, cytochrome c acts as an electron carrier in the electron transport chain, facilitating adenosine triphosphate synthesis, regulating cardiolipin peroxidation, and influencing reactive oxygen species dynamics. Upon cellular stress, it can be released into the cytosol, where it interacts with apoptotic peptidase activator 1 (APAF1) to form the apoptosome, initiating caspase-dependent apoptotic cell death. Additionally, following exposure to pro-apoptotic compounds, cytochrome c contributes to the survival of drug-tolerant persister cells. When translocated to the nucleus, it can induce chromatin condensation and disrupt nucleosome assembly. Upon its release into the extracellular space, cytochrome c may act as an immune mediator during cell death processes, highlighting its multifaceted role in cellular biology. In this review, we explore the diverse structural and functional aspects of cytochrome c in physiological and pathological responses. We summarize how posttranslational modifications of cytochrome c (e.g., phosphorylation, acetylation, tyrosine nitration, and oxidation), binding proteins (e.g., HIGD1A, CHCHD2, ITPR1, and nucleophosmin), and mutations (e.g., G41S, Y48H, and A51V) affect its function. Furthermore, we provide an overview of the latest advanced technologies utilized for detecting cytochrome c, along with potential therapeutic approaches related to this protein. These strategies hold tremendous promise in personalized health care, presenting opportunities for targeted interventions in a wide range of conditions, including neurodegenerative disorders, cardiovascular diseases, and cancer.
© 2024. The Author(s), under exclusive licence to ADMC Associazione Differenziamento e Morte Cellulare.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Tissue-specific regulation of cytochrome c by post-translational modifications: respiration, the mitochondrial membrane potential, ROS, and apoptosis.FASEB J. 2019 Feb;33(2):1540-1553. doi: 10.1096/fj.201801417R. Epub 2018 Sep 17. FASEB J. 2019. PMID: 30222078 Free PMC article. Review.
-
Apoptosis and human diseases: mitochondrion damage and lethal role of released cytochrome C as proapoptotic protein.Curr Med Chem. 2009;16(31):4058-65. doi: 10.2174/092986709789378206. Curr Med Chem. 2009. PMID: 19754424 Review.
-
Copper metabolism in cell death and autophagy.Autophagy. 2023 Aug;19(8):2175-2195. doi: 10.1080/15548627.2023.2200554. Epub 2023 Apr 16. Autophagy. 2023. PMID: 37055935 Free PMC article. Review.
-
Comparison of the structural dynamic and mitochondrial electron-transfer properties of the proapoptotic human cytochrome c variants, G41S, Y48H and A51V.J Inorg Biochem. 2020 Feb;203:110924. doi: 10.1016/j.jinorgbio.2019.110924. Epub 2019 Nov 14. J Inorg Biochem. 2020. PMID: 31760234
-
Apoptotic interactions of cytochrome c: redox flirting with anionic phospholipids within and outside of mitochondria.Biochim Biophys Acta. 2006 May-Jun;1757(5-6):648-59. doi: 10.1016/j.bbabio.2006.03.002. Epub 2006 Mar 31. Biochim Biophys Acta. 2006. PMID: 16740248 Review.
Cited by
-
In Silico Analysis of Temperature-Induced Structural, Stability, and Flexibility Modulations in Camel Cytochrome c.Animals (Basel). 2025 Jan 28;15(3):381. doi: 10.3390/ani15030381. Animals (Basel). 2025. PMID: 39943151 Free PMC article.
-
Selective apoptosis induction by antimicrobial peptide TP4 in MCF-7 breast cancer cells.Sci Rep. 2025 Apr 29;15(1):15061. doi: 10.1038/s41598-025-97906-3. Sci Rep. 2025. PMID: 40301390 Free PMC article.
-
Mitochondrial Calcium Uniporter (MCU)-Mediated Calcium Overload in Psychoactive Drug Neurotoxicity: From Pathogenesis to Therapeutic Targets.Int J Mol Sci. 2025 May 15;26(10):4732. doi: 10.3390/ijms26104732. Int J Mol Sci. 2025. PMID: 40429873 Free PMC article. Review.
-
Macroautophagy/autophagy promotes resistance to KRASG12D-targeted therapy through glutathione synthesis.Cancer Lett. 2024 Nov 1;604:217258. doi: 10.1016/j.canlet.2024.217258. Epub 2024 Sep 12. Cancer Lett. 2024. PMID: 39276914
-
Parthenolide improves sepsis-induced coagulopathy by inhibiting mitochondrial-mediated apoptosis in vascular endothelial cells through BRD4/BCL-xL pathway.J Transl Med. 2025 Jan 17;23(1):80. doi: 10.1186/s12967-025-06114-0. J Transl Med. 2025. PMID: 39825405 Free PMC article.
References
-
- Munn CAM, Foster MVI. Researches on myohamatin and the histohæmatins. Philos Trans R Soc Lond. 1886;177:267–98.
-
- Keilin D. On cytochrome, a respiratory pigment, common to animals, yeast, and higher plants. Proc R Soc Lond B. 1925;98:312–39. doi: 10.1098/rspb.1925.0039. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

