Prokaryotic DNA Crossroads: Holliday Junction Formation and Resolution
- PMID: 38524412
- PMCID: PMC10956419
- DOI: 10.1021/acsomega.3c09866
Prokaryotic DNA Crossroads: Holliday Junction Formation and Resolution
Abstract
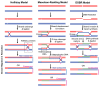
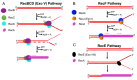
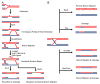
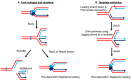
Cells are continually exposed to a multitude of internal and external stressors, which give rise to various types of DNA damage. To protect the integrity of their genetic material, cells are equipped with a repertoire of repair proteins that engage in various repair mechanisms, facilitated by intricate networks of protein-protein and protein-DNA interactions. Among these networks is the homologous recombination (HR) system, a molecular repair mechanism conserved in all three domains of life. On one hand, HR ensures high-fidelity, template-dependent DNA repair, while on the other hand, it results in the generation of combinatorial genetic variations through allelic exchange. Despite substantial progress in understanding this pathway in bacteria, yeast, and humans, several critical questions remain unanswered, including the molecular processes leading to the exchange of DNA segments, the coordination of protein binding, conformational switching during branch migration, and the resolution of Holliday Junctions (HJs). This Review delves into our current understanding of the HR pathway in bacteria, shedding light on the roles played by various proteins or their complexes at different stages of HR. In the first part of this Review, we provide a brief overview of the end resection processes and the strand-exchange reaction, offering a concise depiction of the mechanisms that culminate in the formation of HJs. In the latter half, we expound upon the alternative methods of branch migration and HJ resolution more comprehensively and holistically, considering the historical research timelines. Finally, when we consolidate our knowledge about HR within the broader context of genome replication and the emergence of resistant species, it becomes evident that the HR pathway is indispensable for the survival of bacteria in diverse ecological niches.
© 2024 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
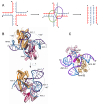
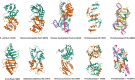
References
-
- Muller H. J. The gene. Proc. R Soc. Med. 1947, 134, 1–37. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
