Molecular Property Diagnostic Suite for COVID-19 (MPDSCOVID-19): an open-source disease-specific drug discovery portal
- PMID: 38525218
- PMCID: PMC10958779
- DOI: 10.46471/gigabyte.114
Molecular Property Diagnostic Suite for COVID-19 (MPDSCOVID-19): an open-source disease-specific drug discovery portal
Abstract
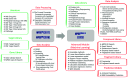
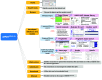
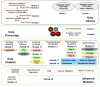
Molecular Property Diagnostic Suite (MPDS) was conceived and developed as an open-source disease-specific web portal based on Galaxy. MPDSCOVID-19 was developed for COVID-19 as a one-stop solution for drug discovery research. Galaxy platforms enable the creation of customized workflows connecting various modules in the web server. The architecture of MPDSCOVID-19 effectively employs Galaxy v22.04 features, which are ported on CentOS 7.8 and Python 3.7. MPDSCOVID-19 provides significant updates and the addition of several new tools updated after six years. Tools developed by our group in Perl/Python and open-source tools are collated and integrated into MPDSCOVID-19 using XML scripts. Our MPDS suite aims to facilitate transparent and open innovation. This approach significantly helps bring inclusiveness in the community while promoting free access and participation in software development.
Availability & implementation: The MPDSCOVID-19 portal can be accessed at https://mpds.neist.res.in:8085/.
© The Author(s) 2024.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Galaxy for open-source computational drug discovery solutions.Expert Opin Drug Discov. 2023 Jun;18(6):579-590. doi: 10.1080/17460441.2023.2205122. Epub 2023 Apr 23. Expert Opin Drug Discov. 2023. PMID: 37089036
-
Molecular property diagnostic suite (MPDS): Development of disease-specific open source web portals for drug discovery.SAR QSAR Environ Res. 2017 Nov;28(11):913-926. doi: 10.1080/1062936X.2017.1402819. SAR QSAR Environ Res. 2017. PMID: 29206500
-
Molecular Property Diagnostic Suite Compound Library (MPDS-CL): a structure-based classification of the chemical space.Mol Divers. 2024 Oct;28(5):3243-3259. doi: 10.1007/s11030-023-10752-1. Epub 2023 Oct 30. Mol Divers. 2024. PMID: 37902900
-
INSPIRE datahub: a pan-African integrated suite of services for harmonising longitudinal population health data using OHDSI tools.Front Digit Health. 2024 Jan 29;6:1329630. doi: 10.3389/fdgth.2024.1329630. eCollection 2024. Front Digit Health. 2024. PMID: 38347885 Free PMC article. Review.
-
Open-Source Browser-Based Tools for Structure-Based Computer-Aided Drug Discovery.Molecules. 2022 Jul 20;27(14):4623. doi: 10.3390/molecules27144623. Molecules. 2022. PMID: 35889494 Free PMC article. Review.
Cited by
-
Pharmacoinformatics in identifying therapeutically important chemical species from Ayurvedic formulations employed in treating COVID-19 patients.J Ayurveda Integr Med. 2025 Jul-Aug;16(4):101161. doi: 10.1016/j.jaim.2025.101161. Epub 2025 Jun 23. J Ayurveda Integr Med. 2025. PMID: 40555008 Free PMC article.
References
LinkOut - more resources
Full Text Sources
