Quorum-sensing regulation of phenazine production heightens Pseudomonas aeruginosa resistance to ciprofloxacin
- PMID: 38526048
- PMCID: PMC11064481
- DOI: 10.1128/aac.00118-24
Quorum-sensing regulation of phenazine production heightens Pseudomonas aeruginosa resistance to ciprofloxacin
Abstract
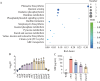
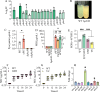
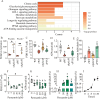
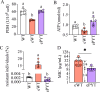
Quorum sensing is a type of cell-cell communication that modulates various biological activities of bacteria. Previous studies indicate that quorum sensing contributes to the evolution of bacterial resistance to antibiotics, but the underlying mechanisms are not fully understood. In this study, we grew Pseudomonas aeruginosa in the presence of sub-lethal concentrations of ciprofloxacin, resulting in a large increase in ciprofloxacin minimal inhibitory concentration. We discovered that quorum sensing-mediated phenazine biosynthesis was significantly enhanced in the resistant isolates, where the quinolone circuit was the predominant contributor to this phenomenon. We found that production of pyocyanin changed carbon flux and showed that the effect can be partially inhibited by the addition of pyruvate to cultures. This study illustrates the role of quorum sensing-mediated phenotypic resistance and suggests a strategy for its prevention.
Keywords: Pseudomonas aeruginosa; antimicrobial resistance; pyocyanin; quorum sensing.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Differential Regulation of the Phenazine Biosynthetic Operons by Quorum Sensing in Pseudomonas aeruginosa PAO1-N.Front Cell Infect Microbiol. 2018 Jul 23;8:252. doi: 10.3389/fcimb.2018.00252. eCollection 2018. Front Cell Infect Microbiol. 2018. PMID: 30083519 Free PMC article.
-
Expression of the las and rhl quorum-sensing systems in clinical isolates of Pseudomonas aeruginosa does not correlate with efflux pump expression or antimicrobial resistance.J Antimicrob Chemother. 2006 Dec;58(6):1250-3. doi: 10.1093/jac/dkl407. Epub 2006 Oct 9. J Antimicrob Chemother. 2006. PMID: 17030516
-
Ostarine attenuates pyocyanin in Pseudomonas aeruginosa by interfering with quorum sensing systems.J Antibiot (Tokyo). 2021 Dec;74(12):863-873. doi: 10.1038/s41429-021-00469-4. Epub 2021 Sep 3. J Antibiot (Tokyo). 2021. PMID: 34480092
-
The third quorum-sensing system of Pseudomonas aeruginosa: Pseudomonas quinolone signal and the enigmatic PqsE protein.J Med Microbiol. 2020 Jan;69(1):25-34. doi: 10.1099/jmm.0.001116. J Med Microbiol. 2020. PMID: 31794380 Review.
-
Marine natural products as potential anti-Pseudomonas aeruginosa agents: challenges and advances.Eur J Med Chem. 2025 Aug 5;292:117670. doi: 10.1016/j.ejmech.2025.117670. Epub 2025 Apr 21. Eur J Med Chem. 2025. PMID: 40305937 Review.
Cited by
-
Characterization of AI-2/LuxS quorum sensing system in antibiotic resistance, pathogenicity of non carbapenemase-producing carbapenem-resistant Escherichia coli.BMC Microbiol. 2025 Mar 15;25(1):140. doi: 10.1186/s12866-025-03846-3. BMC Microbiol. 2025. PMID: 40087577 Free PMC article.
References
-
- O’Neil J. 2014. Antimicrobial resistance: tackling a crisis for the health and wealth of nations. In Rev Antimicrob resist. Vol. 1.
-
- Antimicrobial resistance. Available from: www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance
-
- Eklöf J, Sørensen R, Ingebrigtsen TS, Sivapalan P, Achir I, Boel JB, Bangsborg J, Ostergaard C, Dessau RB, Jensen US, Browatzki A, Lapperre TS, Janner J, Weinreich UM, Armbruster K, Wilcke T, Seersholm N, Jensen JUS. 2020. Pseudomonas aeruginosa and risk of death and exacerbations in patients with chronic obstructive pulmonary disease: an observational cohort study of 22053 patients. Clin Microbiol Infect 26:227–234. doi:10.1016/j.cmi.2019.06.011 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

