Lipopolysaccharide Regulates the Macrophage RNA-Binding Proteome
- PMID: 38527097
- PMCID: PMC11296930
- DOI: 10.1021/acs.jproteome.3c00838
Lipopolysaccharide Regulates the Macrophage RNA-Binding Proteome
Abstract
RNA-protein interactions within cellular signaling pathways have significant modulatory effects on RNA binding proteins' (RBPs') effector functions. During the innate immune response, specific RNA-protein interactions have been reported as a regulatory layer of post-transcriptional control. We investigated changes in the RNA-bound proteome of immortalized mouse macrophages (IMM) following treatment with lipopolysaccharide (LPS). Stable isotope labeling by amino acids in cell culture (SILAC) of cells followed by unbiased purification of RNP complexes at two time points after LPS stimulation and bottom-up proteomic analysis by LC-MS/MS resulted in a set of significantly affected RBPs. Global RNA sequencing and LFQ proteomics were used to characterize the correlation of transcript and protein abundance changes in response to LPS at different time points with changes in protein-RNA binding. Il1α, MARCKS, and ACOD1 were noted as RBP candidates involved in innate immune signaling. The binding sites of the RBP and RNA conjugates at amino acid resolution were investigated by digesting the cross-linked oligonucleotide from peptides remaining after elution using Nuclease P1. The combined data sets provide directions for further studies of innate immune signaling regulation by RBP interactions with different classes of RNA.
Keywords: RNA-binding proteins; TLR signaling; cell signaling; innate immunity; macrophages; protein−RNA interactions; proteomics.
Conflict of interest statement
Conflicts of Interest
The authors declare no conflict of interest.
Figures



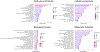


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

