Extend the benchmarking indel set by manual review using the individual cell line sequencing data from the Sequencing Quality Control 2 (SEQC2) project
- PMID: 38528062
- PMCID: PMC10963753
- DOI: 10.1038/s41598-024-57439-7
Extend the benchmarking indel set by manual review using the individual cell line sequencing data from the Sequencing Quality Control 2 (SEQC2) project
Abstract
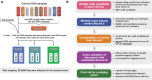
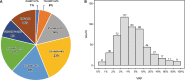
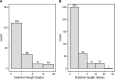
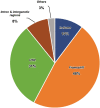
Accurate indel calling plays an important role in precision medicine. A benchmarking indel set is essential for thoroughly evaluating the indel calling performance of bioinformatics pipelines. A reference sample with a set of known-positive variants was developed in the FDA-led Sequencing Quality Control Phase 2 (SEQC2) project, but the known indels in the known-positive set were limited. This project sought to provide an enriched set of known indels that would be more translationally relevant by focusing on additional cancer related regions. A thorough manual review process completed by 42 reviewers, two advisors, and a judging panel of three researchers significantly enriched the known indel set by an additional 516 indels. The extended benchmarking indel set has a large range of variant allele frequencies (VAFs), with 87% of them having a VAF below 20% in reference Sample A. The reference Sample A and the indel set can be used for comprehensive benchmarking of indel calling across a wider range of VAF values in the lower range. Indel length was also variable, but the majority were under 10 base pairs (bps). Most of the indels were within coding regions, with the remainder in the gene regulatory regions. Although high confidence can be derived from the robust study design and meticulous human review, this extensive indel set has not undergone orthogonal validation. The extended benchmarking indel set, along with the indels in the previously published known-positive set, was the truth set used to benchmark indel calling pipelines in a community challenge hosted on the precisionFDA platform. This benchmarking indel set and reference samples can be utilized for a comprehensive evaluation of indel calling pipelines. Additionally, the insights and solutions obtained during the manual review process can aid in improving the performance of these pipelines.
Keywords: Benchmarking; Bioinformatics; Indel; Precision medicine; Quality control.
© 2024. This is a U.S. Government work and not under copyright protection in the US; foreign copyright protection may apply.
Conflict of interest statement
N.F.S. and N.N. are employees of Agilent Technologies. Other authors declare no competing interest.
Figures
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

