Development of Genome-Wide Intron Length Polymorphism (ILP) Markers in Tea Plant (Camellia sinensis) and Related Applications for Genetics Research
- PMID: 38542215
- PMCID: PMC10969888
- DOI: 10.3390/ijms25063241
Development of Genome-Wide Intron Length Polymorphism (ILP) Markers in Tea Plant (Camellia sinensis) and Related Applications for Genetics Research
Abstract
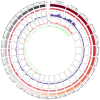
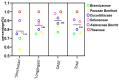
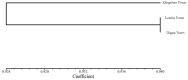
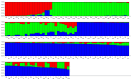
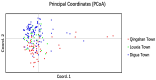
The market value of tea is largely dependent on the tea species and cultivar. Therefore, it is important to develop efficient molecular markers covering the entire tea genome that can be used for the identification of tea varieties, marker-assisted breeding, and mapping important quantitative trait loci for beneficial traits. In this study, genome-wide molecular markers based on intron length polymorphism (ILP) were developed for tea trees. A total of 479, 1393, and 1342 tea ILP markers were identified using the PCR method in silico from the 'Shuchazao' scaffold genome, the chromosome-level genome of 'Longjing 43', and the ancient tea DASZ chromosome-level genome, respectively. A total of 230 tea ILP markers were used to amplify six tea tree species. Among these, 213 pairs of primers successfully characterize products in all six species, with 112 primer pairs exhibiting polymorphism. The polymorphism rate of primer pairs increased with the improvement in reference genome assembly quality level. The cross-species transferability analysis of 35 primer pairs of tea ILP markers showed an average amplification rate of 85.17% through 11 species in 6 families, with high transferability in Camellia reticulata and tobacco. We also used 40 pairs of tea ILP primers to evaluate the genetic diversity and population structure of C. tetracocca with 176 plants from Puan County, Guizhou Province, China. These genome-wide markers will be a valuable resource for genetic diversity analysis, marker-assisted breeding, and variety identification in tea, providing important information for the tea industry.
Keywords: Camellia tetracocca; genetic diversity; intron length polymorphism (ILP); population structure; transferability.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

Similar articles
-
Genome-wide identification and development of miniature inverted-repeat transposable elements and intron length polymorphic markers in tea plant (Camellia sinensis).Sci Rep. 2022 Sep 28;12(1):16233. doi: 10.1038/s41598-022-20400-7. Sci Rep. 2022. PMID: 36171247 Free PMC article.
-
Genome-wide development and deployment of informative intron-spanning and intron-length polymorphism markers for genomics-assisted breeding applications in chickpea.Plant Sci. 2016 Nov;252:374-387. doi: 10.1016/j.plantsci.2016.08.013. Epub 2016 Aug 25. Plant Sci. 2016. PMID: 27717474
-
Characterization of genome-wide genetic variations between two varieties of tea plant (Camellia sinensis) and development of InDel markers for genetic research.BMC Genomics. 2019 Dec 5;20(1):935. doi: 10.1186/s12864-019-6347-0. BMC Genomics. 2019. PMID: 31805860 Free PMC article.
-
Understanding the Origin and Evolution of Tea (Camellia sinensis [L.]): Genomic Advances in Tea.J Mol Evol. 2023 Apr;91(2):156-168. doi: 10.1007/s00239-023-10099-z. Epub 2023 Mar 1. J Mol Evol. 2023. PMID: 36859501 Review.
-
Application of Multi-Perspectives in Tea Breeding and the Main Directions.Int J Mol Sci. 2023 Aug 10;24(16):12643. doi: 10.3390/ijms241612643. Int J Mol Sci. 2023. PMID: 37628823 Free PMC article. Review.
Cited by
-
Genetic Diversity and Population Structure of Wild Ancient Camellia tetracocca in Pu'an, Guizhou, China.Plants (Basel). 2025 Jun 4;14(11):1709. doi: 10.3390/plants14111709. Plants (Basel). 2025. PMID: 40508383 Free PMC article.
-
Application of an Anchor Mapping of Alien Chromosome (AMAC) Fragment Localization Method in the Identification of Radish Chromosome Segments in the Progeny of Rape-Radish Interspecific Hybrids.Int J Mol Sci. 2024 Dec 21;25(24):13687. doi: 10.3390/ijms252413687. Int J Mol Sci. 2024. PMID: 39769448 Free PMC article.
-
Increasing jute (Corchorus olitorius L.) fiber yield through hybridization and combining ability studies to break the yield plateau.Front Plant Sci. 2025 Feb 26;16:1499256. doi: 10.3389/fpls.2025.1499256. eCollection 2025. Front Plant Sci. 2025. PMID: 40078629 Free PMC article.
-
Rapid Identification of Alien Chromosome Fragments and Tracing of Bioactive Compound Genes in Intergeneric Hybrid Offspring Between Brassica napus and Isatis indigotica Based on AMAC Method.Int J Mol Sci. 2025 Feb 27;26(5):2091. doi: 10.3390/ijms26052091. Int J Mol Sci. 2025. PMID: 40076717 Free PMC article.
References
-
- Chen L., Yao M.Z., Wang X.C., Yang Y.J. Tea genetic resources in China. Int. J. Tea Sci. 2012;8:55–64.
-
- Yao M.-Z., Ma C.-L., Qiao T.-T., Jin J.-Q., Chen L. Diversity distribution and population structure of tea germplasms in China revealed by EST-SSR markers. Tree Genet. Genomes. 2012;8:205–220. doi: 10.1007/s11295-011-0433-z. - DOI
MeSH terms
Substances
Grants and funding
- YNWR-QNBJ-2019-280 and YNWR-QNBJ-2020-222/the Outstanding Young Talents Support Program of Yunnan Province
- 202101BD070001-041/the Agricultural Joint Special Project - General Project of Yunnan
- 202305AD160025/Yunnan Program of Technology Innovation and Talent Cultivation
- 202205AR070001/Yunnan Seed laboratory
- 202101BD070001-095/Yunnan agricultural basic research joint special general project
LinkOut - more resources
Full Text Sources

