Inhibitor Trapping in Kinases
- PMID: 38542228
- PMCID: PMC10970472
- DOI: 10.3390/ijms25063249
Inhibitor Trapping in Kinases
Abstract
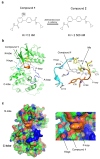
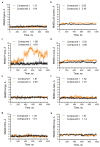
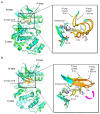
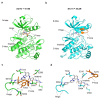
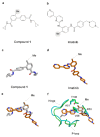
Recently, we identified a novel mechanism of enzyme inhibition in N-myristoyltransferases (NMTs), which we have named 'inhibitor trapping'. Inhibitor trapping occurs when the protein captures the small molecule within its structural confines, thereby preventing its free dissociation and resulting in a dramatic increase in inhibitor affinity and potency. Here, we demonstrate that inhibitor trapping also occurs in the kinases. Remarkably, the drug imatinib, which has revolutionized targeted cancer therapy, is entrapped in the structure of the Abl kinase. This effect is also observed in p38α kinase, where inhibitor trapping was found to depend on a 'magic' methyl group, which stabilizes the protein conformation and increases the affinity of the compound dramatically. Altogether, these results suggest that inhibitor trapping is not exclusive to N-myristoyltransferases, as it also occurs in the kinase family. Inhibitor trapping could enhance the binding affinity of an inhibitor by thousands of times and is as a key mechanism that plays a critical role in determining drug affinity and potency.
Keywords: Abl; Src; binding affinity; binding affinity prediction; drug design; drug potency; imatinib; kinases; protein conformation.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
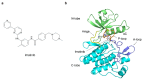




References
-
- Dian C., Perez-Dorado I., Riviere F., Asensio T., Legrand P., Ritzefeld M., Shen M., Cota E., Meinnel T., Tate E.W., et al. High-resolution snapshots of human N-myristoyltransferase in action illuminate a mechanism promoting N-terminal Lys and Gly myristoylation. Nat. Commun. 2020;11:1132. doi: 10.1038/s41467-020-14847-3. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

