Whole-Genome Survey Analyses of Five Goby Species Provide Insights into Their Genetic Evolution and Invasion-Related Genes
- PMID: 38542267
- PMCID: PMC10970681
- DOI: 10.3390/ijms25063293
Whole-Genome Survey Analyses of Five Goby Species Provide Insights into Their Genetic Evolution and Invasion-Related Genes
Abstract
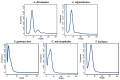
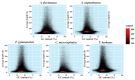
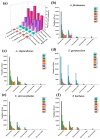
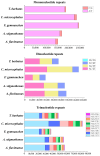
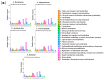
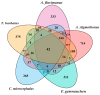
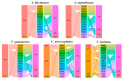
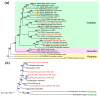
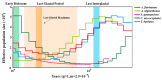
As one of the most abundant groups in marine fish families, Gobiidae fish are important fishery resources in China, and some are also invasive species in certain regions worldwide. However, the phylogenetic relationships of Gobiidae fish remain ambiguous, and the study of their invasion-related genes is still scarce. This study used high-throughput sequencing technology to conduct a whole-genome survey of five Gobiidae fish species: Acanthogobius flavimanus, Acanthogobius stigmothonus, Favonigobius gymnauchen, Ctenotrypauchen microcephalus, and Tridentiger barbatus. De novo assembly of five fish genomes was performed, and genomic traits were compared through K-mer analysis. Among the five Gobiidae fish genomes, F. gymnauchen had the largest genome size (1601.98 Mb) and the highest heterozygosity (1.56%) and repeat rates (59.83%). Phylogenetic studies showed that A. flavimanus was most closely linked to A. stigmothonus, while Apogonidae and Gobiidae were closely related families. PSMC analysis revealed that C. microcephalus experienced a notable population expansion than the other four fish species in the Early Holocene. By using the KOG, GO, and KEGG databases to annotate single-copy genes, the annotated genes of the five fish were mainly classified as "signal transduction mechanisms", "cellular process", "cellular anatomical entity", and "translation". Acanthogobius flavimanus, A. stigmothonus, and T. barbatus had more genes classified as "response to stimulus" and "localization", which may have played an important role in their invasive processes. Our study also provides valuable material about Gobiidae fish genomics and genetic evolution.
Keywords: Gobiidae; PSMC; microsatellite; phylogenetic; whole-genome survey.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


Similar articles
-
Complete mitochondrial genome sequence of Tridentiger bifasciatus and Tridentiger barbatus (Perciformes, Gobiidae): a mitogenomic perspective on the phylogenetic relationships of Gobiidae.Mol Biol Rep. 2015 Jan;42(1):253-65. doi: 10.1007/s11033-014-3768-3. Epub 2014 Sep 27. Mol Biol Rep. 2015. PMID: 25260906
-
The complete mitochondrial genome of Acanthogobius stigmothonus (Perciformes, Gobiidae).Mitochondrial DNA B Resour. 2020 Jul 20;5(3):2888-2889. doi: 10.1080/23802359.2020.1791004. Mitochondrial DNA B Resour. 2020. PMID: 33457989 Free PMC article.
-
The complete mitochondrial genome sequence and gene organization of Tridentiger trigonocephalus (Gobiidae: Gobionellinae) with phylogenetic consideration.Mitochondrial DNA A DNA Mapp Seq Anal. 2016 Sep;27(5):3725-6. doi: 10.3109/19401736.2015.1079876. Epub 2015 Sep 15. Mitochondrial DNA A DNA Mapp Seq Anal. 2016. PMID: 26370266
-
The mitochondrial genome sequences of the round goby and the sand goby reveal patterns of recent evolution in gobiid fish.BMC Genomics. 2017 Feb 16;18(1):177. doi: 10.1186/s12864-017-3550-8. BMC Genomics. 2017. PMID: 28209125 Free PMC article.
-
Complete mitochondrial genome of freshwater goby Rhinogobius cliffordpopei (Perciformes, Gobiidae): genome characterization and phylogenetic analysis.Genes Genomics. 2018 Nov;40(11):1137-1148. doi: 10.1007/s13258-018-0669-1. Epub 2018 Feb 15. Genes Genomics. 2018. PMID: 30315517
References
-
- Winterbottom R., Emery A.R. A new genus and two new species of gobiid fishes (Perciformes) from the Chagos Archipelago, Central Indian Ocean. Environ. Biol. Fishes. 1981;6:139–149. doi: 10.1007/BF00002777. - DOI
-
- Froese R. FishBase. World Wide Web Electronic Publication. 2005. [(accessed on 2 January 2024)]. Available online: http://www.fishbase.org.
-
- Han D., Xue Y., Ji Y., Xu B., Liu H., Ma Q. Trophic and spatial niche of five gobiid fishes in Jiaozhou Bay. J. Fish. Sci. China. 2013;20:148–156. doi: 10.3724/SP.J.1118.2013.00148. - DOI
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

