Assessment of Gastroenteric Viruses in Marketed Bivalve Mollusks in the Tourist Cities of Rio de Janeiro, Brazil, 2022
- PMID: 38543684
- PMCID: PMC10974528
- DOI: 10.3390/v16030317
Assessment of Gastroenteric Viruses in Marketed Bivalve Mollusks in the Tourist Cities of Rio de Janeiro, Brazil, 2022
Abstract
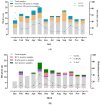
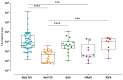
This study investigated the prevalence and genetic diversity of gastroenteric viruses in mussels and oysters in Rio de Janeiro, Brazil. One hundred and thirty-four marketed bivalve samples were obtained between January and December 2022. The viral analysis was performed according to ISO/TS 15216, and the screening revealed the detection of norovirus GII/GI (40.3%), sapovirus (SaV; 12.7%), human mastadenovirus (7.5%), and rotavirus A (RVA; 5.9%). In total, 44.8% (60) of shellfish samples tested positive for one or more viruses, 46.7% (28/60) of the positive samples tested positive for a single viral agent, 26.7% (16) tested positive for two viral agents, 8.3% (5) for three viral agents, and 13.3% (8) for four viral agents. Additionally, three mussel samples were contaminated with the five investigated viruses (5%, 3/60). Norovirus GII showed the highest mean viral load (3.4 × 105 GC/g), followed by SaV (1.4 × 104 GC/g), RVA (1.1 × 104 GC/g), human mastadenovirus (3.9 × 103 GC/g), and norovirus GI (6.7 × 102 GC/g). Molecular characterization revealed that the recovered norovirus strains belonged to genotypes GII.2, GII.6, GII.9, GII.17, and GII.27; SaV belonged to genotypes GI.1 and GIV.1; RVA to genotypes G6, G8, P[8]-III, and human mastadenovirus to types F40 and F41. The GII.27 norovirus characterized in this study is the only strain of this genotype reported in Brazil. This study highlights the dissemination and diversity of gastroenteric viruses present in commercialized bivalves in a touristic area, indicating the potential risk to human health and the contribution of bivalves in the propagation of emerging pathogens.
Keywords: ISO 15216; bivalve mollusk; food safety; gastroenteric viruses; genotyping; monitoring.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



Similar articles
-
Occurrence and Molecular Characterization of Human Astrovirus and Hepatitis A Virus in Bivalve Mollusks Marketed in Tourist Cities in Rio de Janeiro, Brazil.Food Environ Virol. 2025 Apr 2;17(2):23. doi: 10.1007/s12560-025-09639-1. Food Environ Virol. 2025. PMID: 40172833
-
CrAssphage as a Human Enteric Viral Contamination Bioindicator in Marketed Bivalve Mollusks.Viruses. 2025 Jul 18;17(7):1012. doi: 10.3390/v17071012. Viruses. 2025. PMID: 40733628 Free PMC article.
-
Detection and Molecular Characterization of Enteric Viruses in Bivalve Mollusks Collected in Arraial do Cabo, Rio de Janeiro, Brazil.Viruses. 2022 Oct 26;14(11):2359. doi: 10.3390/v14112359. Viruses. 2022. PMID: 36366459 Free PMC article.
-
Monitoring and Genotyping of Norovirus in Bivalve Molluscan Shellfish from Northern Italian Seas (2018-2020).Foodborne Pathog Dis. 2024 Jan;21(1):27-35. doi: 10.1089/fpd.2023.0078. Epub 2023 Oct 25. Foodborne Pathog Dis. 2024. PMID: 37878812
-
Molecular epidemiology and genotype distributions of noroviruses and sapoviruses in Thailand 2000-2016: A review.J Med Virol. 2018 Apr;90(4):617-624. doi: 10.1002/jmv.25019. Epub 2018 Jan 23. J Med Virol. 2018. PMID: 29315631 Review.
Cited by
-
Occurrence and Molecular Characterization of Human Astrovirus and Hepatitis A Virus in Bivalve Mollusks Marketed in Tourist Cities in Rio de Janeiro, Brazil.Food Environ Virol. 2025 Apr 2;17(2):23. doi: 10.1007/s12560-025-09639-1. Food Environ Virol. 2025. PMID: 40172833
-
CrAssphage as a Human Enteric Viral Contamination Bioindicator in Marketed Bivalve Mollusks.Viruses. 2025 Jul 18;17(7):1012. doi: 10.3390/v17071012. Viruses. 2025. PMID: 40733628 Free PMC article.
-
Detection of Rocahepevirus ratti in Bivalve Mollusks from São Luís Island, Maranhão, Brazil: A Potential Transmission Route of an Emerging Zoonotic Pathogen?Food Environ Virol. 2025 Jan 4;17(1):11. doi: 10.1007/s12560-024-09624-0. Food Environ Virol. 2025. PMID: 39754637
-
Human norovirus in Brazil: an update of reports in different settings.Braz J Microbiol. 2024 Sep;55(3):2767-2782. doi: 10.1007/s42770-024-01444-5. Epub 2024 Jul 16. Braz J Microbiol. 2024. PMID: 39012425 Free PMC article. Review.
References
-
- Betts R., de Blackburn C.W. 2-Detecting pathogens in food. In: Blackburn C.d.W., McClure P.J., editors. Food Science, Technology and Nutrition, Foodborne Pathogens. 2nd ed. Woodhead Publishing; Sawston, UK: 2009. pp. 17–65. Woodhead Publishing Series. - DOI
-
- Amaral M.S., Estevam G.K., Penatti M., Lafontaine R., Lima I.C., Spada P.K., Gabbay Y.B., Matos N.B. The prevalence of norovirus, astrovirus and adenovirus infections among hospitalised children with acute gastroenteritis in Porto Velho, state of Rondônia, western Brazilian Amazon. Mem. Inst. Oswaldo Cruz. 2015;110:215–221. doi: 10.1590/0074-02760140381. - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

