The Effect of Global Spread, Epidemiology, and Control Strategies on the Evolution of the GI-19 Lineage of Infectious Bronchitis Virus
- PMID: 38543846
- PMCID: PMC10974917
- DOI: 10.3390/v16030481
The Effect of Global Spread, Epidemiology, and Control Strategies on the Evolution of the GI-19 Lineage of Infectious Bronchitis Virus
Abstract
The GI-19 lineage of infectious bronchitis virus (IBV) has emerged as one of the most impactful, particularly in the "Old World". Originating in China several decades ago, it has consistently spread and evolved, often forming independent clades in various areas and countries, each with distinct production systems and control strategies. This study leverages this scenario to explore how different environments may influence virus evolution. Through the analysis of the complete S1 sequence, four datasets were identified, comprising strains of monophyletic clades circulating in different continents or countries (e.g., Asia vs. Europe and China vs. Thailand), indicative of single introduction events and independent evolution. The population dynamics and evolutionary rate variation over time, as well as the presence and intensity of selective pressures, were estimated and compared across these datasets. Since the lineage origin (approximately in the mid-20th century), a more persistent and stable viral population was estimated in Asia and China, while in Europe and Thailand, a sharp increase following the introduction (i.e., 2005 and 2007, respectively) of GI-19 was observed, succeeded by a rapid decline. Although a greater number of sites on the S1 subunit were under diversifying selection in the Asian and Chinese datasets, more focused and stronger pressures were evident in both the European (positions 2, 52, 54, 222, and 379 and Thai (i.e., positions 10, 12, 32, 56, 62, 64, 65, 78, 95, 96, 119, 128, 140, 182, 292, 304, 320, and 323) strains, likely reflecting a more intense and uniform application of vaccines in these regions. This evidence, along with the analysis of control strategies implemented in different areas, suggests a strong link between effective, systematic vaccine implementation and infection control. However, while the overall evolutionary rate was estimated at approximately 10-3 to 10-4, a significant inverse correlation was found between viral population size and the rate of viral evolution over time. Therefore, despite the stronger selective pressure imposed by vaccination, effectively constraining the former through adequate control strategies can efficiently prevent viral evolution and the emergence of vaccine-escaping variants.
Keywords: IBV; environments; evolution; immunity; natural selection; phylodynamic; vaccine.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

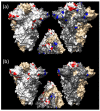
Similar articles
-
Think globally, act locally: Phylodynamic reconstruction of infectious bronchitis virus (IBV) QX genotype (GI-19 lineage) reveals different population dynamics and spreading patterns when evaluated on different epidemiological scales.PLoS One. 2017 Sep 7;12(9):e0184401. doi: 10.1371/journal.pone.0184401. eCollection 2017. PLoS One. 2017. PMID: 28880958 Free PMC article.
-
Evolution and Epidemic Spread of the Avian Infectious Bronchitis Virus (IBV) GI-23 in Brazil.Viruses. 2023 May 24;15(6):1229. doi: 10.3390/v15061229. Viruses. 2023. PMID: 37376528 Free PMC article.
-
Genotyping and phylogeography of infectious bronchitis virus isolates from Pakistan show unique linkage to GI-24 lineage.Poult Sci. 2024 Jan;103(1):103236. doi: 10.1016/j.psj.2023.103236. Epub 2023 Oct 24. Poult Sci. 2024. PMID: 37980750 Free PMC article.
-
The emergence, evolution and spread of infectious bronchitis virus genotype GI-23.Arch Virol. 2021 Jan;166(1):9-26. doi: 10.1007/s00705-020-04920-z. Epub 2021 Jan 8. Arch Virol. 2021. PMID: 33416996 Free PMC article. Review.
-
Severe acute respiratory syndrome vaccine development: experiences of vaccination against avian infectious bronchitis coronavirus.Avian Pathol. 2003 Dec;32(6):567-82. doi: 10.1080/03079450310001621198. Avian Pathol. 2003. PMID: 14676007 Free PMC article. Review.
Cited by
-
Reconstruction of Avian Reovirus History and Dispersal Patterns: A Phylodynamic Study.Viruses. 2024 May 16;16(5):796. doi: 10.3390/v16050796. Viruses. 2024. PMID: 38793677 Free PMC article.
-
Evaluation of Different Machine Learning Approaches to Predict Antigenic Distance Among Newcastle Disease Virus (NDV) Strains.Viruses. 2025 Apr 14;17(4):567. doi: 10.3390/v17040567. Viruses. 2025. PMID: 40285009 Free PMC article.
-
Phylodynamic reconstruction of major chicken infectious anemia virus clades epidemiology, dispersal, and evolution.Front Microbiol. 2025 Jan 17;16:1527335. doi: 10.3389/fmicb.2025.1527335. eCollection 2025. Front Microbiol. 2025. PMID: 39896436 Free PMC article.
-
Evolution and transmission dynamics of infectious bronchitis virus in southern China during 2018-2023: Emergence of a novel GI-19 subtype and re-emergence of GI-28.Poult Sci. 2025 Jul 11;104(10):105564. doi: 10.1016/j.psj.2025.105564. Online ahead of print. Poult Sci. 2025. PMID: 40774171 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

