Expanding the molecular language of protein liquid-liquid phase separation
- PMID: 38553587
- PMCID: PMC11230844
- DOI: 10.1038/s41557-024-01489-x
Expanding the molecular language of protein liquid-liquid phase separation
Abstract
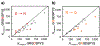
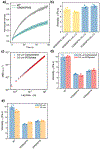
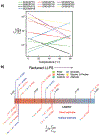
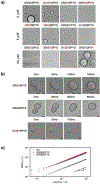
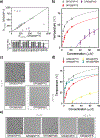
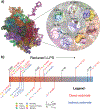
Understanding the relationship between a polypeptide sequence and its phase separation has important implications for analysing cellular function, treating disease and designing novel biomaterials. Several sequence features have been identified as drivers for protein liquid-liquid phase separation (LLPS), schematized as a 'molecular grammar' for LLPS. Here we further probe how sequence modulates phase separation and the material properties of the resulting condensates, targeting sequence features previously overlooked in the literature. We generate sequence variants of a repeat polypeptide with either no charged residues, high net charge, no glycine residues or devoid of aromatic or arginine residues. All but one of 12 variants exhibited LLPS, albeit to different extents, despite substantial differences in composition. Furthermore, we find that all the condensates formed behaved like viscous fluids, despite large differences in their viscosities. Our results support the model of multiple interactions between diverse residue pairs-not just a handful of residues-working in tandem to drive the phase separation and dynamics of condensates.
© 2024. The Author(s), under exclusive licence to Springer Nature Limited.
Conflict of interest statement
Competing Interests
The authors declare no competing interests.
Figures


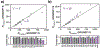





References
-
- Brangwynne CP et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009). - PubMed
-
- Alberti S & Dormann D Liquid–liquid phase separation in disease. Annual review of genetics 53, 171–194 (2019). - PubMed
-
- Alberti S & Hyman AA Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nature reviews Molecular cell biology 22, 196–213 (2021). - PubMed
MeSH terms
Substances
Grants and funding
- DMR-2004796/National Science Foundation (NSF)
- R35 GM153388/GM/NIGMS NIH HHS/United States
- R01GM136917/U.S. Department of Health & Human Services | NIH | Office of Extramural Research, National Institutes of Health (OER)
- R01 GM136917/GM/NIGMS NIH HHS/United States
- R35 GM142903/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Research Materials

