The effects of antibiotic exposures on the gut resistome during hematopoietic cell transplantation in children
- PMID: 38555499
- PMCID: PMC10984140
- DOI: 10.1080/19490976.2024.2333748
The effects of antibiotic exposures on the gut resistome during hematopoietic cell transplantation in children
Abstract
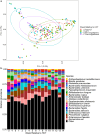
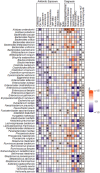
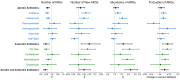
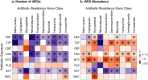
Antibiotic resistance is a global threat driven primarily by antibiotic use. We evaluated the effects of antibiotic exposures on the gut microbiomes and resistomes of children at high risk of colonization by antibiotic-resistant bacteria. We performed shotgun metagenomic sequencing of 691 serially collected fecal samples from 80 children (<18 years) undergoing hematopoietic cell transplantation. We evaluated the effects of aerobic (cefepime, vancomycin, fluoroquinolones, aminoglycosides, macrolides, and trimethoprim-sulfamethoxazole) and anaerobic (piperacillin-tazobactam, carbapenems, metronidazole, and clindamycin) antibiotic exposures on the diversity and composition of the gut microbiome and resistome. We identified 372 unique antibiotic resistance genes (ARGs); the most frequent ARGs identified encode resistance to tetracyclines (n = 88), beta-lactams (n = 84), and fluoroquinolones (n = 79). Both aerobic and anaerobic antibiotic exposures were associated with a decrease in the number of bacterial species (aerobic, β = 0.71, 95% CI: 0.64, 0.79; anaerobic, β = 0.66, 95% CI: 0.53, 0.82) and the number of unique ARGs (aerobic, β = 0.81, 95% CI: 0.74, 0.90; anaerobic, β = 0.73, 95% CI: 0.61, 0.88) within the gut metagenome. However, only antibiotic regimens that included anaerobic activity were associated with an increase in acquisition of new ARGs (anaerobic, β = 1.50; 95% CI: 1.12, 2.01) and an increase in the relative abundance of ARGs in the gut resistome (anaerobic, β = 1.62; 95% CI: 1.15, 2.27). Specific antibiotic exposures were associated with distinct changes in the number and abundance of ARGs for individual antibiotic classes. Our findings detail the impact of antibiotics on the gut microbiome and resistome and demonstrate that anaerobic antibiotics are particularly likely to promote acquisition and expansion of antibiotic-resistant bacteria.
Keywords: Shotgun metagenomic sequencing; anaerobic bacteria; antibiotic resistance; antimicrobial stewardship; carbapenem; metronidazole; piperacillin-tazobactam.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures

Similar articles
-
The infant gut resistome is associated with E. coli and early-life exposures.BMC Microbiol. 2021 Jul 2;21(1):201. doi: 10.1186/s12866-021-02129-x. BMC Microbiol. 2021. PMID: 34215179 Free PMC article.
-
Impact of Antibiotic Treatment on the Gut Microbiome and its Resistome in Hematopoietic Stem Cell Transplant Recipients.J Infect Dis. 2023 Jun 28;228(1):28-36. doi: 10.1093/infdis/jiad033. J Infect Dis. 2023. PMID: 36751730
-
Exposure to doxycycline increases risk of carrying a broad range of enteric antimicrobial resistance determinants in an elderly cohort.J Infect. 2024 Oct;89(4):106243. doi: 10.1016/j.jinf.2024.106243. Epub 2024 Aug 12. J Infect. 2024. PMID: 39142392
-
Global landscape of gut microbiome diversity and antibiotic resistomes across vertebrates.Sci Total Environ. 2022 Sep 10;838(Pt 2):156178. doi: 10.1016/j.scitotenv.2022.156178. Epub 2022 May 23. Sci Total Environ. 2022. PMID: 35618126 Review.
-
Good microbes, bad genes? The dissemination of antimicrobial resistance in the human microbiome.Gut Microbes. 2022 Jan-Dec;14(1):2055944. doi: 10.1080/19490976.2022.2055944. Gut Microbes. 2022. PMID: 35332832 Free PMC article. Review.
Cited by
-
Antibiotic-related dysbiosis of gut microbiota in healthy volunteers: a comparative study of ceftazidime-avibactam, piperacillin-tazobactam, and ceftriaxone.Eur J Clin Microbiol Infect Dis. 2025 Jun 12. doi: 10.1007/s10096-025-05184-8. Online ahead of print. Eur J Clin Microbiol Infect Dis. 2025. PMID: 40504338
-
Management of febrile neutropenia: consensus of the Brazilian Association of Hematology, Blood Transfusion and Cell Therapy - ABHH.Hematol Transfus Cell Ther. 2024 Dec;46 Suppl 6(Suppl 6):S346-S361. doi: 10.1016/j.htct.2024.11.119. Epub 2024 Dec 18. Hematol Transfus Cell Ther. 2024. PMID: 39694764 Free PMC article.
-
The Microbiome and Pediatric Transplantation.J Pediatric Infect Dis Soc. 2024 Feb 28;13(Supplement_1):S80-S89. doi: 10.1093/jpids/piad062. J Pediatric Infect Dis Soc. 2024. PMID: 38417089 Free PMC article. Review.
References
-
- Antibiotic Resistance Threats in the U.S., 2019 (U.S. Department of Health and Human Services, CDC) . 2019.
-
- Nelson RE, Hatfield KM, Wolford H, Samore MH, Scott RD, Reddy SC, Olubajo B, Paul P, Jernigan JA, Baggs J. et al. National estimates of healthcare costs associated with multidrug-resistant bacterial infections among hospitalized patients in the United States. Clin Infect Dis. Jan 29 2021;72(Suppl Supplement_1):S17–15. doi:10.1093/cid/ciaa1581. - DOI - PMC - PubMed
-
- World Health O . Global action plan on antimicrobial resistance. 2015.
-
- Scheich S, Lindner S, Koenig R, Reinheimer C, Wichelhaus TA, Hogardt M, Besier S, Kempf VAJ, Kessel J, Martin H. et al. Clinical impact of colonization with multidrug-resistant organisms on outcome after allogeneic stem cell transplantation in patients with acute myeloid leukemia. Cancer. Jan 15 2018;124(2):286–296. doi:10.1002/cncr.31045. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
