Development of an orally bioavailable mSWI/SNF ATPase degrader and acquired mechanisms of resistance in prostate cancer
- PMID: 38557192
- PMCID: PMC11009648
- DOI: 10.1073/pnas.2322563121
Development of an orally bioavailable mSWI/SNF ATPase degrader and acquired mechanisms of resistance in prostate cancer
Abstract
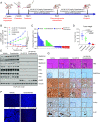
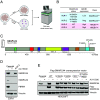
Mammalian switch/sucrose nonfermentable (mSWI/SNF) ATPase degraders have been shown to be effective in enhancer-driven cancers by functioning to impede oncogenic transcription factor chromatin accessibility. Here, we developed AU-24118, an orally bioavailable proteolysis-targeting chimera (PROTAC) degrader of mSWI/SNF ATPases (SMARCA2 and SMARCA4) and PBRM1. AU-24118 demonstrated tumor regression in a model of castration-resistant prostate cancer (CRPC) which was further enhanced with combination enzalutamide treatment, a standard of care androgen receptor (AR) antagonist used in CRPC patients. Importantly, AU-24118 exhibited favorable pharmacokinetic profiles in preclinical analyses in mice and rats, and further toxicity testing in mice showed a favorable safety profile. As acquired resistance is common with targeted cancer therapeutics, experiments were designed to explore potential mechanisms of resistance that may arise with long-term mSWI/SNF ATPase PROTAC treatment. Prostate cancer cell lines exposed to long-term treatment with high doses of a mSWI/SNF ATPase degrader developed SMARCA4 bromodomain mutations and ABCB1 (ATP binding cassette subfamily B member 1) overexpression as acquired mechanisms of resistance. Intriguingly, while SMARCA4 mutations provided specific resistance to mSWI/SNF degraders, ABCB1 overexpression provided broader resistance to other potent PROTAC degraders targeting bromodomain-containing protein 4 and AR. The ABCB1 inhibitor, zosuquidar, reversed resistance to all three PROTAC degraders tested. Combined, these findings position mSWI/SNF degraders for clinical translation for patients with enhancer-driven cancers and define strategies to overcome resistance mechanisms that may arise.
Keywords: ABCB1; SMARCA2; SMARCA4; mSWI/SNF; proteolysis-targeting chimera (PROTAC).
Conflict of interest statement
Competing interests statement:A.M.C. serves on the clinical advisory board of Aurigene Oncology Limited. B.K., S. Mukherjee, S.D., K.B.A., S.D.S., and C.A. are employees of Aurigene Oncology Limited. Aurigene has filed patent applications on AU-15330 and AU-24118. The other authors declare no competing interests.
Figures


Update of
-
Development of an orally bioavailable mSWI/SNF ATPase degrader and acquired mechanisms of resistance in prostate cancer.bioRxiv [Preprint]. 2024 Mar 2:2024.02.29.582768. doi: 10.1101/2024.02.29.582768. bioRxiv. 2024. Update in: Proc Natl Acad Sci U S A. 2024 Apr 9;121(15):e2322563121. doi: 10.1073/pnas.2322563121. PMID: 38464081 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

