Broad-spectrum Delta-BA.2 tandem-fused heterodimer mRNA vaccine delivered by lipopolyplex
- PMID: 38557908
- PMCID: PMC11008869
- DOI: 10.1371/journal.ppat.1012116
Broad-spectrum Delta-BA.2 tandem-fused heterodimer mRNA vaccine delivered by lipopolyplex
Abstract
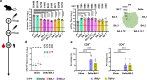
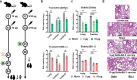
Since the beginning of the coronavirus disease 2019 (COVID-19) pandemic, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative agent of COVID-19, continues to mutate and generates new variants with increasingly severe immune escape, urging the upgrade of COVID-19 vaccines. Here, based on a similar dimeric RBD design as our previous ZF2001 vaccine, we developed a novel broad-spectrum COVID-19 mRNA vaccine, SWIM516, with chimeric Delta-BA.2 RBD dimer delivered by lipopolyplex (LPP). Unlike the popular lipid nanoparticle (LNP), this LPP-delivered mRNA expresses only in the injection site, which avoids potential toxicity to the liver. We demonstrated the broad-spectrum humoral and cellular immunogenicity of this vaccine to Delta and Omicron sub-variants in naïve mice and as booster shots. When challenged with Delta or Omicron live virus, vaccinated human angiotensin-converting enzyme (hACE2) transgenic mice and rhesus macaques were both protected, displaying significantly reduced viral loads and markedly relieved pathological damages. We believe the SWIM516 vaccine qualifies as a candidate for the next-generation broad-spectrum COVID-19 vaccine.
Copyright: © 2024 Du et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

