B-cell-directed CAR T-cell therapy activates CD8+ cytotoxic CARneg bystander T cells in patients and nonhuman primates
- PMID: 38558106
- PMCID: PMC11561542
- DOI: 10.1182/blood.2023022717
B-cell-directed CAR T-cell therapy activates CD8+ cytotoxic CARneg bystander T cells in patients and nonhuman primates
Abstract
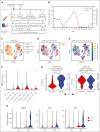
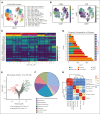
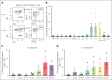
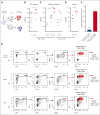
Chimeric antigen receptor (CAR) T cells hold promise as a therapy for B-cell-derived malignancies, and despite their impressive initial response rates, a significant proportion of patients ultimately experience relapse. Although recent studies have explored the mechanisms of in vivo CAR T-cell function, little is understood about the activation of surrounding CARneg bystander T cells and their potential to enhance tumor responses. We performed single-cell RNA sequencing on nonhuman primate (NHP) and patient-derived T cells to identify the phenotypic and transcriptomic hallmarks of bystander activation of CARneg T cells following B-cell-targeted CAR T-cell therapy. Using a highly translatable CD20 CAR NHP model, we observed a distinct population of activated CD8+ CARneg T cells emerging during CAR T-cell expansion. These bystander CD8+ CARneg T cells exhibited a unique transcriptional signature with upregulation of natural killer-cell markers (KIR3DL2, CD160, and KLRD1), chemokines, and chemokine receptors (CCL5, XCL1, and CCR9), and downregulation of naïve T-cell-associated genes (SELL and CD28). A transcriptionally similar population was identified in patients after a tisagenlecleucel infusion. Mechanistic studies revealed that interleukin-2 (IL-2) and IL-15 exposure induced bystander-like CD8+ T cells in a dose-dependent manner. In vitro activated and patient-derived T cells with a bystander phenotype efficiently killed leukemic cells through a T-cell receptor-independent mechanism. Collectively, to our knowledge, these data provide the first comprehensive identification and profiling of CARneg bystander CD8+ T cells following B-cell-targeting CAR T-cell therapy and suggest a novel mechanism through which CAR T-cell infusion might trigger enhanced antileukemic responses. Patient samples were obtained from the trial #NCT03369353, registered at www.ClinicalTrials.gov.
© 2024 American Society of Hematology. Published by Elsevier Inc. All rights are reserved, including those for text and data mining, AI training, and similar technologies.
Conflict of interest statement
Conflict-of-interest disclosure: U.G. possesses intellectual property rights related to AlloVir, including interests in royalties. L.S.K. is on the scientific advisory board of Mammoth Biosciences and HiFiBIO; reports research funding from Magenta Therapeutics, Tessera Therapeutics, Novartis, Emmanuel Merck, Darmstadt Serono, Gilead Pharmaceuticals, and Regeneron Pharmaceuticals; reports consulting fees from Vertex; reports grants and personal fees from Bristol Myers Squibb; and the conflict-of-interest between L.S.K. with Bristol Myers Squibb is managed under an agreement with the Harvard Medical School. A.K.S. reports compensation for consulting and/or scientific advisory board membership from Merck, Honeycomb Biotechnologies, Cellarity, Repertoire Immune Medicines, Ochre Bio, Third Rock Ventures, Hovione, Relation Therapeutics, FL82, FL86, Empress Therapeutics, IntrECate Biotherapeutics, Senda Biosciences, and Dahlia Biosciences unrelated to this work. Parts of the study were supported by 2seventy bio. F.E. and E.E.H. are employees of 2seventy bio, J.B.R. is an employee of Tessara therapeutics, and S.P.M. is an employee of Cue Biopharma. The remaining authors declare no competing financial interests.
Figures



References
-
- Majzner RG, Mackall CL. Clinical lessons learned from the first leg of the CAR T cell journey. Nat Med. 2019;25(9):1341–1355. - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

