Genome-wide identification of tea plant (Camellia sinensis) BAHD acyltransferases reveals their role in response to herbivorous pests
- PMID: 38561653
- PMCID: PMC10985903
- DOI: 10.1186/s12870-024-04867-2
Genome-wide identification of tea plant (Camellia sinensis) BAHD acyltransferases reveals their role in response to herbivorous pests
Abstract
Background: BAHD acyltransferases are among the largest metabolic protein domain families in the genomes of terrestrial plants and play important roles in plant growth and development, aroma formation, and biotic and abiotic stress responses. Little is known about the BAHDs in the tea plant, a cash crop rich in secondary metabolites.
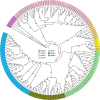
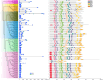
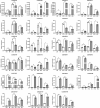
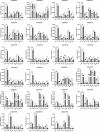
Results: In this study, 112 BAHD genes (CsBAHD01-CsBAHD112) were identified from the tea plant genome, with 85% (98/112) unevenly distributed across the 15 chromosomes. The number of BAHD gene family members has significantly expanded from wild tea plants to the assamica type to the sinensis type. Phylogenetic analysis showed that they could be classified into seven subgroups. Promoter cis-acting element analysis revealed that they contain a large number of light, phytohormones, and stress-responsive elements. Many members displayed tissue-specific expression patterns. CsBAHD05 was expressed at more than 500-fold higher levels in purple tea leaves than in green tea leaves. The genes exhibiting the most significant response to MeJA treatment and feeding by herbivorous pests were primarily concentrated in subgroups 5 and 6. The expression of 23 members of these two subgroups at different time points after feeding by tea green leafhoppers and tea geometrids was examined via qPCR, and the results revealed that the expression of CsBAHD93, CsBAHD94 and CsBAHD95 was significantly induced after the tea plants were subjected to feeding by both pricking and chewing pests. Moreover, based on the transcriptome data for tea plants being fed on by these two pests, a transcriptional regulatory network of different transcription factor genes coexpressed with these 23 members was constructed.
Conclusions: Our study provides new insights into the role of BAHDs in the defense response of tea plants, and will facilitate in-depth studies of the molecular function of BAHDs in resistance to herbivorous pests.
Keywords: Camellia sinensis; Acyltransferase; HIPVs; Tea geometrid; Tea green leafhopper.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Genome-wide identification and expression analysis of SABATH methyltransferases in tea plant (Camellia sinensis): insights into their roles in plant defense responses.Plant Signal Behav. 2020 Oct 2;15(10):1804684. doi: 10.1080/15592324.2020.1804684. Epub 2020 Aug 12. Plant Signal Behav. 2020. PMID: 32787495 Free PMC article.
-
A defensive pathway from NAC and TCP transcription factors activates a BAHD acyltransferase for (Z)-3-hexenyl acetate biosynthesis to resist herbivore in tea plant (Camellia sinensis).New Phytol. 2025 Feb;245(3):1232-1248. doi: 10.1111/nph.20283. Epub 2024 Nov 17. New Phytol. 2025. PMID: 39550628
-
Genome-wide identification and expression characterization of the GH3 gene family of tea plant (Camellia sinensis).BMC Genomics. 2024 Jan 27;25(1):120. doi: 10.1186/s12864-024-10004-y. BMC Genomics. 2024. PMID: 38280985 Free PMC article.
-
Biosynthesis and the Transcriptional Regulation of Terpenoids in Tea Plants (Camellia sinensis).Int J Mol Sci. 2023 Apr 8;24(8):6937. doi: 10.3390/ijms24086937. Int J Mol Sci. 2023. PMID: 37108101 Free PMC article. Review.
-
Advances in research on functional genes of tea plant.Gene. 2019 Aug 30;711:143940. doi: 10.1016/j.gene.2019.143940. Epub 2019 Jun 18. Gene. 2019. PMID: 31226279 Review.
Cited by
-
Hydroxycinnamoyl-coenzyme A: tetrahydroxyhexanedioate hydroxycinnamoyl transferase (HHHT) from Phaseolus vulgaris L.: phylogeny, expression pattern, kinetic parameters, and active site analysis.PeerJ. 2025 Feb 20;13:e19037. doi: 10.7717/peerj.19037. eCollection 2025. PeerJ. 2025. PMID: 39989742 Free PMC article.
-
Plant Defense Proteins: Recent Discoveries and Applications.Plants (Basel). 2025 Jul 6;14(13):2069. doi: 10.3390/plants14132069. Plants (Basel). 2025. PMID: 40648078 Free PMC article. Review.
-
Genome-wide identification and expression analysis of the BAHD gene family in Leonurus japonicus.Front Genet. 2024 Dec 19;15:1512692. doi: 10.3389/fgene.2024.1512692. eCollection 2024. Front Genet. 2024. PMID: 39748946 Free PMC article.
-
Genome-Wide Identification and Functional Characterization of the BAHD Acyltransferase Gene Family in Brassica napus L.Plants (Basel). 2025 Jul 15;14(14):2183. doi: 10.3390/plants14142183. Plants (Basel). 2025. PMID: 40733420 Free PMC article.
References
-
- St-Pierre B, De Luca V. Chapter nine - evolution of acyltransferase genes: origin and diversification of the BAHD superfamily of acyltransferases involved in secondary metabolism. In: Recent Advances in Phytochemistry Edited by Romeo JT, Ibrahim R, Varin L, De Luca V, vol. 34: Elsevier; 2000: 285–315.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

