This is a preprint.
Ether lipids influence cancer cell fate by modulating iron uptake
- PMID: 38562716
- PMCID: PMC10983928
- DOI: 10.1101/2024.03.20.585922
Ether lipids influence cancer cell fate by modulating iron uptake
Abstract
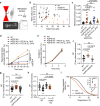
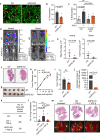
Cancer cell fate has been widely ascribed to mutational changes within protein-coding genes associated with tumor suppressors and oncogenes. In contrast, the mechanisms through which the biophysical properties of membrane lipids influence cancer cell survival, dedifferentiation and metastasis have received little scrutiny. Here, we report that cancer cells endowed with a high metastatic ability and cancer stem cell-like traits employ ether lipids to maintain low membrane tension and high membrane fluidity. Using genetic approaches and lipid reconstitution assays, we show that these ether lipid-regulated biophysical properties permit non-clathrin-mediated iron endocytosis via CD44, leading directly to significant increases in intracellular redox-active iron and enhanced ferroptosis susceptibility. Using a combination of in vitro three-dimensional microvascular network systems and in vivo animal models, we show that loss of ether lipids also strongly attenuates extravasation, metastatic burden and cancer stemness. These findings illuminate a mechanism whereby ether lipids in carcinoma cells serve as key regulators of malignant progression while conferring a unique vulnerability that can be exploited for therapeutic intervention.
Keywords: CD44; Ether lipids; endocytosis; ferroptosis; iron; membrane tension; metastasis.
Conflict of interest statement
COMPETING INTERESTS The authors declare no competing interests.
Figures



Similar articles
-
Activation of lysosomal iron triggers ferroptosis in cancer.Res Sq [Preprint]. 2024 Apr 8:rs.3.rs-4165774. doi: 10.21203/rs.3.rs-4165774/v1. Res Sq. 2024. Update in: Nature. 2025 Jun;642(8067):492-500. doi: 10.1038/s41586-025-08974-4. PMID: 38659936 Free PMC article. Updated. Preprint.
-
A tale of two lipids: Lipid unsaturation commands ferroptosis sensitivity.Proteomics. 2023 Mar;23(6):e2100308. doi: 10.1002/pmic.202100308. Epub 2023 Jan 13. Proteomics. 2023. PMID: 36398995 Review.
-
A novel EHD1/CD44/Hippo/SP1 positive feedback loop potentiates stemness and metastasis in lung adenocarcinoma.Clin Transl Med. 2022 Apr;12(4):e836. doi: 10.1002/ctm2.836. Clin Transl Med. 2022. PMID: 35485206 Free PMC article.
-
Therapy-induced lipid uptake and remodeling underpin ferroptosis hypersensitivity in prostate cancer.Cancer Metab. 2020 Jun 19;8:11. doi: 10.1186/s40170-020-00217-6. eCollection 2020. Cancer Metab. 2020. PMID: 32577235 Free PMC article.
-
Ether Lipids in Obesity: From Cells to Population Studies.Front Physiol. 2022 Mar 3;13:841278. doi: 10.3389/fphys.2022.841278. eCollection 2022. Front Physiol. 2022. PMID: 35309067 Free PMC article. Review.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
