Cultivable microbial diversity, peptide profiles, and bio-functional properties in Parmigiano Reggiano cheese
- PMID: 38567075
- PMCID: PMC10985727
- DOI: 10.3389/fmicb.2024.1342180
Cultivable microbial diversity, peptide profiles, and bio-functional properties in Parmigiano Reggiano cheese
Abstract
Introduction: Lactic acid bacteria (LAB) communities shape the sensorial and functional properties of artisanal hard-cooked and long-ripened cheeses made with raw bovine milk like Parmigiano Reggiano (PR) cheese. While patterns of microbial evolution have been well studied in PR cheese, there is a lack of information about how this microbial diversity affects the metabolic and functional properties of PR cheese.
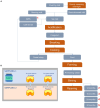
Methods: To fill this information gap, we characterized the cultivable fraction of natural whey starter (NWS) and PR cheeses at different ripening times, both at the species and strain level, and investigated the possible correlation between microbial composition and the evolution of peptide profiles over cheese ripening.
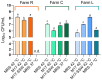
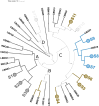
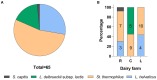
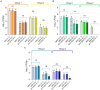
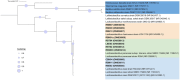
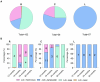
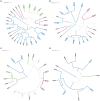
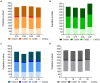
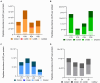
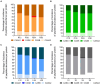
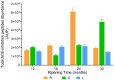
Results and discussion: The results showed that NWS was a complex community of several biotypes belonging to a few species, namely, Streptococcus thermophilus, Lactobacillus helveticus, and Lactobacillus delbrueckii subsp. lactis. A new species-specific PCR assay was successful in discriminating the cheese-associated species Lacticaseibacillus casei, Lacticaseibacillus paracasei, Lacticaseibacillus rhamnosus, and Lacticaseibacillus zeae. Based on the resolved patterns of species and biotype distribution, Lcb. paracasei and Lcb. zeae were most frequently isolated after 24 and 30 months of ripening, while the number of biotypes was inversely related to the ripening time. Peptidomics analysis revealed more than 520 peptides in cheese samples. To the best of our knowledge, this is the most comprehensive survey of peptides in PR cheese. Most of them were from β-caseins, which represent the best substrate for LAB cell-envelope proteases. The abundance of peptides from β-casein 38-88 region continuously increased during ripening. Remarkably, this region contains precursors for the anti-hypertensive lactotripeptides VPP and IPP, as well as for β-casomorphins. We found that the ripening time strongly affects bioactive peptide profiles and that the occurrence of Lcb. zeae species is positively linked to the incidence of eight anti-hypertensive peptides. This result highlighted how the presence of specific LAB species is likely a pivotal factor in determining PR functional properties.
Keywords: Lacticaseibacillus; Parmigiano Reggiano cheese; bioactive peptides; natural whey starter; non-starter lactic acid bacteria; peptidomics; starter lactic acid bacteria.
Copyright © 2024 Martini, Sola, Cattivelli, Cristofolini, Pizzamiglio, Tagliazucchi and Solieri.
Conflict of interest statement
VP was employed by Consorzio del Formaggio Parmigiano Reggiano. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures

Similar articles
-
Cultivable non-starter lactobacilli from ripened Parmigiano Reggiano cheeses with different salt content and their potential to release anti-hypertensive peptides.Int J Food Microbiol. 2020 Oct 2;330:108688. doi: 10.1016/j.ijfoodmicro.2020.108688. Epub 2020 May 29. Int J Food Microbiol. 2020. PMID: 32497940
-
The Interrelationship Between Microbiota and Peptides During Ripening as a Driver for Parmigiano Reggiano Cheese Quality.Front Microbiol. 2020 Oct 2;11:581658. doi: 10.3389/fmicb.2020.581658. eCollection 2020. Front Microbiol. 2020. PMID: 33133050 Free PMC article.
-
Evaluation of genetic polymorphism among Lactobacillus rhamnosus non-starter Parmigiano Reggiano cheese strains.Int J Food Microbiol. 2011 Jan 5;144(3):569-72. doi: 10.1016/j.ijfoodmicro.2010.11.017. Epub 2010 Nov 13. Int J Food Microbiol. 2011. PMID: 21131087
-
The Microfloras and Sensory Profiles of Selected Protected Designation of Origin Italian Cheeses.Microbiol Spectr. 2014 Feb;2(1):CM-0007-2012. doi: 10.1128/microbiolspec.CM-0007-2012. Microbiol Spectr. 2014. PMID: 26082116 Review.
-
Invited review: Microbial evolution in raw-milk, long-ripened cheeses produced using undefined natural whey starters.J Dairy Sci. 2014 Feb;97(2):573-91. doi: 10.3168/jds.2013-7187. Epub 2013 Dec 2. J Dairy Sci. 2014. PMID: 24290824 Review.
References
-
- Adamberg K., Antonsson M., Vogensen F. K., Nielsen E. W., Kask S., Møller P. L., et al. . (2005). Fermentation of carbohydrates from cheese sources by non-starter lactic acid bacteria isolated from semi-hard Danish cheese. Int. Dairy J. 15, 873–882. doi: 10.1016/j.idairyj.2004.07.017 - DOI
-
- Beresford T. P., Fitzsimons N. A., Brennan N. L., Cogan T. M. (2001). Recent advances in cheese microbiology. Int. Dairy J. 11, 259–274. doi: 10.1016/S0958-6946(01)00056-5 - DOI
-
- Bertani G., Levante A., Lazzi C., Bottari B., Gatti M., Neviani E. (2020). Dynamics of a natural bacterial community under technological and environmental pressures: the case of natural whey starter for Parmigiano Reggiano cheese. Food Res. Int. 129:108860. doi: 10.1016/j.foodres.2019.108860, PMID: - DOI - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

