Engineering Plant Cell Fates and Functions for Agriculture and Industry
- PMID: 38573786
- PMCID: PMC11036505
- DOI: 10.1021/acssynbio.4c00047
Engineering Plant Cell Fates and Functions for Agriculture and Industry
Abstract
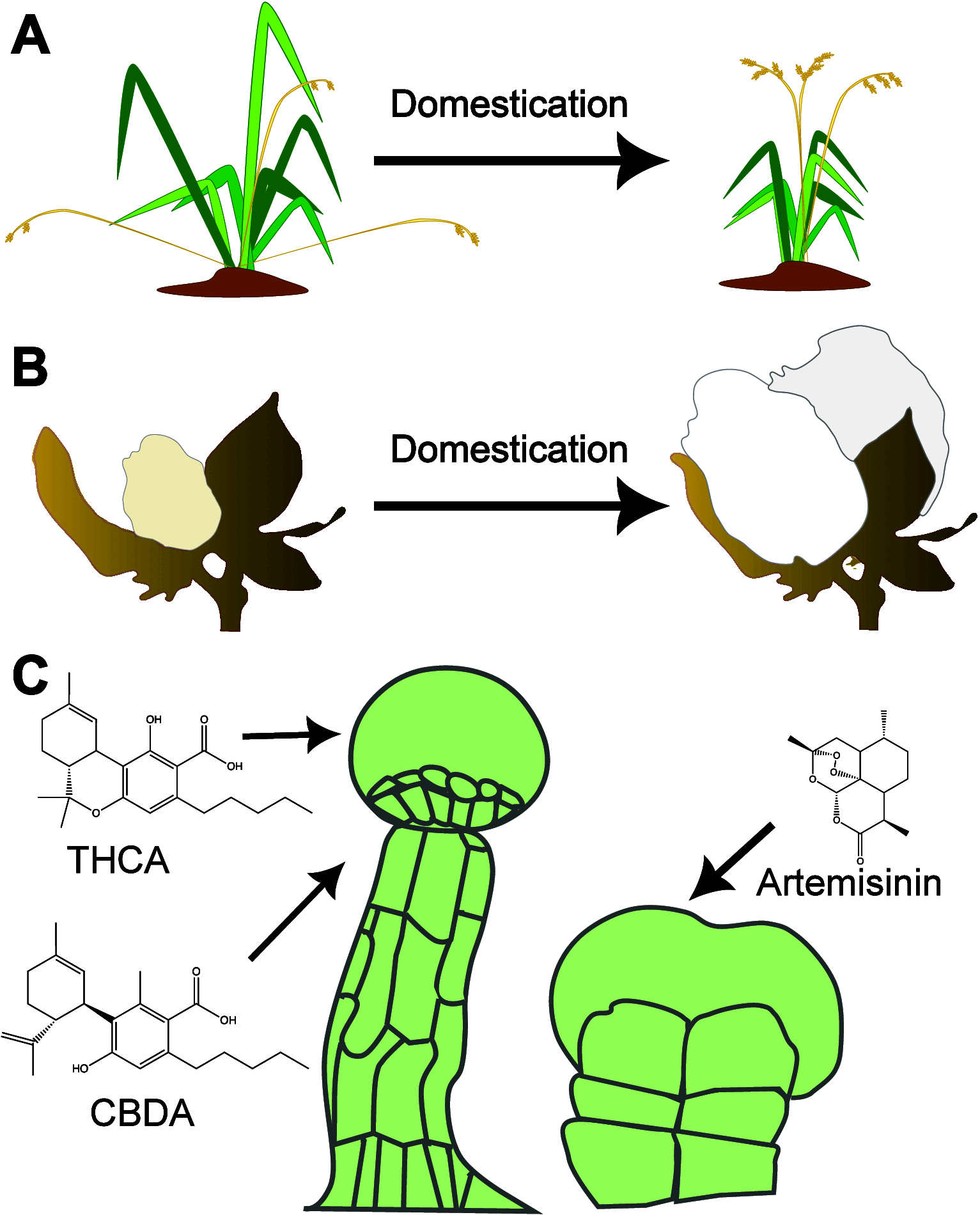
Many plant species are grown to enable access to specific organs or tissues, such as seeds, fruits, or stems. In some cases, a value is associated with a molecule that accumulates in a single type of cell. Domestication and subsequent breeding have often increased the yields of these target products by increasing the size, number, and quality of harvested organs and tissues but also via changes to overall plant growth architecture to suit large-scale cultivation. Many of the mutations that underlie these changes have been identified in key regulators of cellular identity and function. As key determinants of yield, these regulators are key targets for synthetic biology approaches to engineer new forms and functions. However, our understanding of many plant developmental programs and cell-type specific functions is still incomplete. In this Perspective, we discuss how advances in cellular genomics together with synthetic biology tools such as biosensors and DNA-recording devices are advancing our understanding of cell-specific programs and cell fates. We then discuss advances and emerging opportunities for cell-type-specific engineering to optimize plant morphology, responses to the environment, and the production of valuable compounds.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

References
-
- Chandrasekhar A.; Dunne D.; Viglione G.. UN land report: Five key takeaways for climate change, food systems and nature loss. Carbon Brief. April 27, 2022. https://www.carbonbrief.org/un-land-report-five-key-takeaways-for-climat... (accessed 2024–01–15).
-
- Lin T.; Zhu G.; Zhang J.; Xu X.; Yu Q.; Zheng Z.; Zhang Z.; Lun Y.; Li S.; Wang X.; Huang Z.; Li J.; Zhang C.; Wang T.; Zhang Y.; Wang A.; Zhang Y.; Lin K.; Li C.; Xiong G.; Xue Y.; Mazzucato A.; Causse M.; Fei Z.; Giovannoni J. J.; Chetelat R. T.; Zamir D.; Städler T.; Li J.; Ye Z.; Du Y.; Huang S. Genomic Analyses Provide Insights into the History of Tomato Breeding. Nat. Genet. 2014, 46 (11), 1220–1226. 10.1038/ng.3117. - DOI - PubMed
-
- Pnueli L.; Carmel-Goren L.; Hareven D.; Gutfinger T.; Alvarez J.; Ganal M.; Zamir D.; Lifschitz E. The SELF-PRUNING Gene of Tomato Regulates Vegetative to Reproductive Switching of Sympodial Meristems and Is the Ortholog of CEN and TFL1. Development 1998, 125 (11), 1979–1989. 10.1242/dev.125.11.1979. - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

