Oxidative stress is intrinsic to staphylococcal adaptation to fatty acid synthesis antibiotics
- PMID: 38577105
- PMCID: PMC10993138
- DOI: 10.1016/j.isci.2024.109505
Oxidative stress is intrinsic to staphylococcal adaptation to fatty acid synthesis antibiotics
Abstract
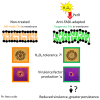
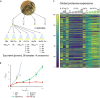
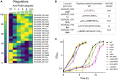
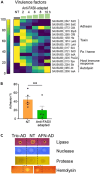
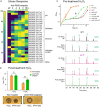
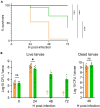
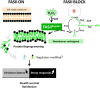
Antibiotics inhibiting the fatty acid synthesis pathway (FASII) of the major pathogen Staphylococcus aureus reach their enzyme targets, but bacteria continue growth by using environmental fatty acids (eFAs) to produce phospholipids. We assessed the consequences and effectors of FASII-antibiotic (anti-FASII) adaptation. Anti-FASII induced lasting expression changes without genomic rearrangements. Several identified regulators affected the timing of adaptation outgrowth. Adaptation resulted in decreased expression of major virulence factors. Conversely, stress responses were globally increased and adapted bacteria were more resistant to peroxide killing. Importantly, pre-exposure to peroxide led to faster anti-FASII-adaptation by stimulating eFA incorporation. This adaptation differs from reports of peroxide-stimulated antibiotic efflux, which leads to tolerance. In vivo, anti-FASII-adapted S. aureus killed the insect host more slowly but continued multiplying. We conclude that staphylococcal adaptation to FASII antibiotics involves reprogramming, which decreases virulence and increases stress resistance. Peroxide, produced by the host to combat infection, favors anti-FASII adaptation.
Keywords: Microbial metabolism; Microbiology.
© 2024 The Authors. Published by Elsevier Inc.
Conflict of interest statement
The authors declare that no competing interests exist.
Figures
Similar articles
-
(p)ppGpp/GTP and Malonyl-CoA Modulate Staphylococcus aureus Adaptation to FASII Antibiotics and Provide a Basis for Synergistic Bi-Therapy.mBio. 2021 Feb 2;12(1):e03193-20. doi: 10.1128/mBio.03193-20. mBio. 2021. PMID: 33531402 Free PMC article.
-
Permissive Fatty Acid Incorporation Promotes Staphylococcal Adaptation to FASII Antibiotics in Host Environments.Cell Rep. 2019 Dec 17;29(12):3974-3982.e4. doi: 10.1016/j.celrep.2019.11.071. Cell Rep. 2019. PMID: 31851927
-
Clinical Relevance of Type II Fatty Acid Synthesis Bypass in Staphylococcus aureus.Antimicrob Agents Chemother. 2017 Apr 24;61(5):e02515-16. doi: 10.1128/AAC.02515-16. Print 2017 May. Antimicrob Agents Chemother. 2017. PMID: 28193654 Free PMC article.
-
The Staphylococcus aureus FASII bypass escape route from FASII inhibitors.Biochimie. 2017 Oct;141:40-46. doi: 10.1016/j.biochi.2017.07.004. Epub 2017 Jul 17. Biochimie. 2017. PMID: 28728970 Review.
-
Mining Fatty Acid Biosynthesis for New Antimicrobials.Annu Rev Microbiol. 2022 Sep 8;76:281-304. doi: 10.1146/annurev-micro-041320-110408. Epub 2022 Jun 1. Annu Rev Microbiol. 2022. PMID: 35650664 Free PMC article. Review.
Cited by
-
Staphylococcus epidermidis uses the SrrAB regulatory system to modulate oxidative stress and intracellular survival in mouse macrophage cell line Ana-1.mSystems. 2025 May 20;10(5):e0173724. doi: 10.1128/msystems.01737-24. Epub 2025 Apr 22. mSystems. 2025. PMID: 40261004 Free PMC article.
References
-
- Cusumano J.A., Klinker K.P., Huttner A., Luther M.K., Roberts J.A., LaPlante K.L. Towards precision medicine: Therapeutic drug monitoring-guided dosing of vancomycin and beta-lactam antibiotics to maximize effectiveness and minimize toxicity. Am. J. Health Syst. Pharm. 2020;77:1104–1112. doi: 10.1093/ajhp/zxaa128. - DOI - PubMed
LinkOut - more resources
Full Text Sources

