14-3-3 binding motif phosphorylation disrupts Hdac4-organized condensates to stimulate cardiac reprogramming
- PMID: 38578832
- PMCID: PMC11081035
- DOI: 10.1016/j.celrep.2024.114054
14-3-3 binding motif phosphorylation disrupts Hdac4-organized condensates to stimulate cardiac reprogramming
Abstract
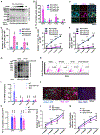
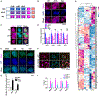
Cell fate conversion is associated with extensive post-translational modifications (PTMs) and architectural changes of sub-organelles, yet how these events are interconnected remains unknown. We report here the identification of a phosphorylation code in 14-3-3 binding motifs (PC14-3-3) that greatly stimulates induced cardiomyocyte (iCM) formation from fibroblasts. PC14-3-3 is identified in pivotal functional proteins for iCM reprogramming, including transcription factors and chromatin modifiers. Akt1 kinase and protein phosphatase 2A are the key writer and key eraser of the PC14-3-3 code, respectively. PC14-3-3 activation induces iCM formation with the presence of only Tbx5. In contrast, PC14-3-3 inhibition by mutagenesis or inhibitor-mediated code removal abolishes reprogramming. We discover that key PC14-3-3-embedded factors, such as histone deacetylase 4 (Hdac4), Mef2c, and Foxo1, form Hdac4-organized inhibitory nuclear condensates. PC14-3-3 activation disrupts Hdac4 condensates to promote cardiac gene expression. Our study suggests that sub-organelle dynamics regulated by a PTM code could be a general mechanism for stimulating cell reprogramming.
Keywords: 14-3-3; CP: Molecular biology; biomolecular condensate; cardiac reprogramming; epigenetic code; post-translational modification.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures



Update of
-
14-3-3 binding motif phosphorylation disrupts Hdac4 organized condensates to stimulate cardiac reprogramming.bioRxiv [Preprint]. 2023 Nov 20:2023.11.20.567913. doi: 10.1101/2023.11.20.567913. bioRxiv. 2023. Update in: Cell Rep. 2024 Apr 23;43(4):114054. doi: 10.1016/j.celrep.2024.114054. PMID: 38045244 Free PMC article. Updated. Preprint.
References
-
- Christoforou N, Chellappan M, Adler AF, Kirkton RD, Wu T, Addis RC, Bursac N, and Leong KW (2013). Transcription factors MYOCD, SRF, Mesp1 and SMARCD3 enhance the cardio-inducing effect of GATA4, TBX5, and MEF2C during direct cellular reprogramming. PLoS One 8, e63577. 10.1371/journal.pone.0063577. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

