Genetic dissection of the tissue-specific roles of type III effectors and phytotoxins in the pathogenicity of Pseudomonas syringae pv. syringae to cherry
- PMID: 38590135
- PMCID: PMC11002349
- DOI: 10.1111/mpp.13451
Genetic dissection of the tissue-specific roles of type III effectors and phytotoxins in the pathogenicity of Pseudomonas syringae pv. syringae to cherry
Abstract
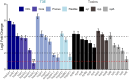
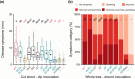
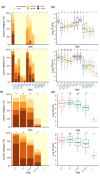
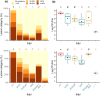
When compared with other phylogroups (PGs) of the Pseudomonas syringae species complex, P. syringae pv. syringae (Pss) strains within PG2 have a reduced repertoire of type III effectors (T3Es) but produce several phytotoxins. Effectors within the cherry pathogen Pss 9644 were grouped based on their frequency in strains from Prunus as the conserved effector locus (CEL) common to most P. syringae pathogens; a core of effectors common to PG2; a set of PRUNUS effectors common to cherry pathogens; and a FLEXIBLE set of T3Es. Pss 9644 also contains gene clusters for biosynthesis of toxins syringomycin, syringopeptin and syringolin A. After confirmation of virulence gene expression, mutants with a sequential series of T3E and toxin deletions were pathogenicity tested on wood, leaves and fruits of sweet cherry (Prunus avium) and leaves of ornamental cherry (Prunus incisa). The toxins had a key role in disease development in fruits but were less important in leaves and wood. An effectorless mutant retained some pathogenicity to fruit but not wood or leaves. Striking redundancy was observed amongst effector groups. The CEL effectors have important roles during the early stages of leaf infection and possibly acted synergistically with toxins in all tissues. Deletion of separate groups of T3Es had more effect in P. incisa than in P. avium. Mixed inocula were used to complement the toxin mutations in trans and indicated that strain mixtures may be important in the field. Our results highlight the niche-specific role of toxins in P. avium tissues and the complexity of effector redundancy in the pathogen Pss 9644.
Keywords: Prunus; Pseudomonas syringae; comparative genomics; mutagenesis; phytotoxins; type 3 effectors; virulence.
© 2024 The Authors. Molecular Plant Pathology published by British Society for Plant Pathology and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Pathogenicity, phylogenomic, and comparative genomic study of Pseudomonas syringae sensu lato affecting sweet cherry in California.Microbiol Spectr. 2024 Oct 3;12(10):e0132424. doi: 10.1128/spectrum.01324-24. Epub 2024 Sep 3. Microbiol Spectr. 2024. PMID: 39225473 Free PMC article.
-
Comparative genomics of Pseudomonas syringae reveals convergent gene gain and loss associated with specialization onto cherry (Prunus avium).New Phytol. 2018 Jul;219(2):672-696. doi: 10.1111/nph.15182. Epub 2018 May 4. New Phytol. 2018. PMID: 29726587
-
Comparative genomics and pathogenicity potential of members of the Pseudomonas syringae species complex on Prunus spp.BMC Genomics. 2019 Mar 5;20(1):172. doi: 10.1186/s12864-019-5555-y. BMC Genomics. 2019. PMID: 30836956 Free PMC article.
-
Pseudomonas syringae pv. syringae Associated With Mango Trees, a Particular Pathogen Within the "Hodgepodge" of the Pseudomonas syringae Complex.Front Plant Sci. 2019 May 8;10:570. doi: 10.3389/fpls.2019.00570. eCollection 2019. Front Plant Sci. 2019. PMID: 31139201 Free PMC article. Review.
-
Closing the circle on the discovery of genes encoding Hrp regulon members and type III secretion system effectors in the genomes of three model Pseudomonas syringae strains.Mol Plant Microbe Interact. 2006 Nov;19(11):1151-8. doi: 10.1094/MPMI-19-1151. Mol Plant Microbe Interact. 2006. PMID: 17073298 Review.
Cited by
-
Pathogenicity, phylogenomic, and comparative genomic study of Pseudomonas syringae sensu lato affecting sweet cherry in California.Microbiol Spectr. 2024 Oct 3;12(10):e0132424. doi: 10.1128/spectrum.01324-24. Epub 2024 Sep 3. Microbiol Spectr. 2024. PMID: 39225473 Free PMC article.
-
A Sucrose-Utilisation Gene Cluster Contributes to Colonisation of Horse Chestnut by Pseudomonas syringae pv. aesculi.Mol Plant Pathol. 2025 Jul;26(7):e70116. doi: 10.1111/mpp.70116. Mol Plant Pathol. 2025. PMID: 40616241 Free PMC article.
References
-
- Alfano, J.R. , Charkowski, A.O. , Deng, W.‐L. , Badel, J.L. , Petnicki‐Ocwieja, T. , Van Dijk, K. et al. (2000) The Pseudomonas syringae Hrp pathogenicity island has a tripartite mosaic structure composed of a cluster of type III secretion genes bounded by exchangeable effector and conserved effector loci that contribute to parasitic fitness and pathogenicity in plants. Proceedings of the National Academy of Sciences of the United States of America, 97, 4856–4861. - PMC - PubMed
-
- Altschul, S.F. , Gish, W. , Miller, W. , Myers, E.W. & Lipman, D.J. (1990) Basic local alignment search tool. Journal of Molecular Biology, 215, 403–410. - PubMed
-
- Andrews, S. (2010) FastQC: a quality control tool for high throughput sequence data. Cambridge, UK: Babraham Bioinformatics, Babraham Institute.
-
- Aronesty, E. (2013) Comparison of sequencing utility programs. The Open Bioinformatics Journal, 7, TOBIOIJ‐7‐1.
-
- Badel, J.L. , Nomura, K. , Bandyopadhyay, S. , Shimizu, R. , Collmer, A. & He, S.Y. (2003) Pseudomonas syringae pv. tomato DC3000 HopPtoM (CEL ORF3) is important for lesion formation but not growth in tomato and is secreted and translocated by the Hrp type III secretion system in a chaperone‐dependent manner. Molecular Microbiology, 49, 1239–1251. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

