Genome-wide identification of bHLH transcription factors and functional analysis in salt gland development of the recretohalophyte sea lavender (Limonium bicolor)
- PMID: 38595909
- PMCID: PMC11001596
- DOI: 10.1093/hr/uhae036
Genome-wide identification of bHLH transcription factors and functional analysis in salt gland development of the recretohalophyte sea lavender (Limonium bicolor)
Abstract
Transcription factors with basic helix-loop-helix (bHLH) structures regulate plant growth, epidermal structure development, metabolic processes, and responses to stress extensively. Sea lavender (Limonium bicolor) is a recretohalophyte with unique salt glands in the epidermis that make it highly resistant to salt stress, contributing to the improvement of saline lands. However, the features of the bHLH transcription factor family in L. bicolor are largely unknown. Here, we systematically analyzed the characteristics, localization, and phylogenetic relationships of 187 identified bHLH family genes throughout the L. bicolor genome, as well as their cis-regulatory promoter elements, expression patterns, and key roles in salt gland development or salt tolerance by genetic analysis. Nine verified L. bicolor bHLH genes are expressed and the encoded proteins function in the nucleus, among which the proteins encoded by Lb2G14060 and Lb1G07934 also localize to salt glands. Analysis of CRISPR-Cas9-generated knockout mutants and overexpression lines indicated that the protein encoded by Lb1G07934 is involved in the formation of salt glands, salt secretion, and salt resistance, indicating that bHLH genes strongly influence epidermal structure development and stress responses. The current study lays the foundation for further investigation of the effects and functional mechanisms of bHLH genes in L. bicolor and paves the way for selecting salt-tolerance genes that will enhance salt resistance in crops and for the improvement of saline soils.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nanjing Agricultural University.
Conflict of interest statement
The authors state that they do not have conflicts of interest related to this work.
Figures


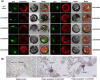


Similar articles
-
The WD40 gene family in recretohalophyte Limonium bicolor: genomic identification and functional analysis in salt gland development and salinity tolerance.Front Plant Sci. 2025 Jul 30;16:1629604. doi: 10.3389/fpls.2025.1629604. eCollection 2025. Front Plant Sci. 2025. PMID: 40810015 Free PMC article.
-
Genome-wide identification of the mitogen-activated kinase gene family from Limonium bicolor and functional characterization of LbMAPK2 under salt stress.BMC Plant Biol. 2023 Nov 15;23(1):565. doi: 10.1186/s12870-023-04589-x. BMC Plant Biol. 2023. PMID: 37964233 Free PMC article.
-
The WRKY gene family in the halophyte Limonium bicolor: identification, expression analysis, and regulation of salt stress tolerance.Plant Cell Rep. 2024 Jun 12;43(7):167. doi: 10.1007/s00299-024-03258-z. Plant Cell Rep. 2024. PMID: 38865016
-
Basic Helix-Loop-Helix Transcription Factors: Regulators for Plant Growth Development and Abiotic Stress Responses.Int J Mol Sci. 2023 Jan 11;24(2):1419. doi: 10.3390/ijms24021419. Int J Mol Sci. 2023. PMID: 36674933 Free PMC article. Review.
-
Progress in Studying Salt Secretion from the Salt Glands in Recretohalophytes: How Do Plants Secrete Salt?Front Plant Sci. 2016 Jun 30;7:977. doi: 10.3389/fpls.2016.00977. eCollection 2016. Front Plant Sci. 2016. PMID: 27446195 Free PMC article. Review.
Cited by
-
Integrated transcriptomic and metabolomic analyses uncover the key pathways of Limonium bicolor in response to salt stress.Plant Biotechnol J. 2025 Mar;23(3):715-730. doi: 10.1111/pbi.14534. Epub 2024 Dec 5. Plant Biotechnol J. 2025. PMID: 39636615 Free PMC article.
-
The WD40 gene family in recretohalophyte Limonium bicolor: genomic identification and functional analysis in salt gland development and salinity tolerance.Front Plant Sci. 2025 Jul 30;16:1629604. doi: 10.3389/fpls.2025.1629604. eCollection 2025. Front Plant Sci. 2025. PMID: 40810015 Free PMC article.
-
Physiological and transcriptomic analysis reveals the coating of microcapsules embedded with bacteria can enhance wheat salt tolerance.BMC Plant Biol. 2024 Oct 25;24(1):1004. doi: 10.1186/s12870-024-05718-w. BMC Plant Biol. 2024. PMID: 39448914 Free PMC article.
References
-
- Yuan J, Zhang S, Liu Z. et al. . Cloning and phylogenetic analysis of an amphioxus myogenic bHLH gene AmphiMDF. Biochem Biophys Res Commun. 2003;301:960–7 - PubMed
-
- Sun W, Xiu J, Ma Z. et al. . Basic helix–loop–helix (bHLH) gene family in Tartary buckwheat (Fagopyrum tataricum): genome-wide identification, phylogeny, evolutionary expansion and expression analyses. Int J Biol Macromol. 2020;155:1478–90 - PubMed
LinkOut - more resources
Full Text Sources

