Oncolytic herpes simplex viruses designed for targeted treatment of EGFR-bearing tumors
- PMID: 38596286
- PMCID: PMC10869753
- DOI: 10.1016/j.omton.2024.200761
Oncolytic herpes simplex viruses designed for targeted treatment of EGFR-bearing tumors
Abstract
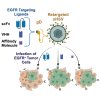
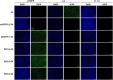
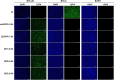
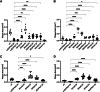
Oncolytic herpes simplex viruses (oHSVs) have emerged as leading cancer therapeutic agents. Effective oHSV virotherapy may ultimately require both intratumoral and systemic vector administration to target the primary tumor and distant metastases. An attractive approach to enhancing oHSV tumor specificity is engineering the virus envelope glycoproteins for selective recognition of and infection via tumor-specific cell surface proteins. We previously demonstrated that oHSVs could be retargeted to EGFR-expressing cells by the incorporation of a single-chain antibody (scFv) at the N terminus of glycoprotein D (gD). Here, we compared retargeted oHSVs generated by the insertion of scFv, affibody molecule, or VHH antibody ligands at different positions within the N terminus of gD. When compared to the scFv-directed oHSVs, VHH and affibody molecules mediated enhanced EGFR-specific tumor cell entry, spread and cell killing in vitro, and enabled long-term tumor-specific virus replication following intravenous delivery in vivo. Moreover, oHSVs retargeted via a VHH ligand reduced tumor growth upon intravenous injection and achieved complete tumor destruction after intratumoral injection. Systemic oHSV delivery is important for the treatment of metastatic disease, and our enhancements in targeted oHSV design are a critical step in creating an effective tumor-specific oHSVs for safe administration via the bloodstream.
Keywords: EGFR; GBM; HSV; MT: Regular Issue; glycoprotein; oncolytic; vector targeting.
© 2024 The Authors.
Conflict of interest statement
J.B.C. and J.C.G. are inventors of intellectual property licensed to Oncorus Inc. (Cambridge, MA). J.C.G. is a consultant to and Chair of the Scientific Advisory Board of Oncorus Inc. The remaining authors declare no competing interests.
Figures


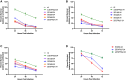
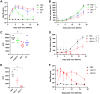
References
-
- Glorioso J.C., Cohen J.B., Goins W.F., Hall B., Jackson J.W., Kohanbash G., Amankulor N., Kaur B., Caligiuri M.A., Chiocca E.A., et al. Oncolytic HSV Vectors and Anti-Tumor Immunity. Curr. Issues Mol. Biol. 2021;41:381–468. - PubMed
-
- Andtbacka R.H.I., Kaufman H.L., Collichio F., Amatruda T., Senzer N., Chesney J., Delman K.A., Spitler L.E., Puzanov I., Agarwala S.S., et al. Talimogene laherparepvec improves durable response rate in patients with advanced melanoma. J. Clin. Oncol. 2015;33:2780–2788. - PubMed
-
- Dambach M.J., Trecki J., Martin N., Markovitz N.S. Oncolytic viruses derived from the gamma34.5-deleted herpes simplex virus recombinant R3616 encode a truncated UL3 protein. Mol. Ther. 2006;13:891–898. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
