Targeting the ribosome to treat multiple myeloma
- PMID: 38596309
- PMCID: PMC10905045
- DOI: 10.1016/j.omton.2024.200771
Targeting the ribosome to treat multiple myeloma
Abstract
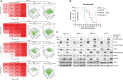
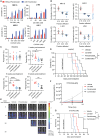
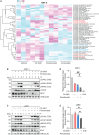
The high rates of protein synthesis and processing render multiple myeloma (MM) cells vulnerable to perturbations in protein homeostasis. The induction of proteotoxic stress by targeting protein degradation with proteasome inhibitors (PIs) has revolutionized the treatment of MM. However, resistance to PIs is inevitable and represents an ongoing clinical challenge. Our first-in-human study of the selective inhibitor of RNA polymerase I transcription of ribosomal RNA genes, CX-5461, has demonstrated a potential signal for anti-tumor activity in three of six heavily pre-treated MM patients. Here, we show that CX-5461 has potent anti-myeloma activity in PI-resistant MM preclinical models in vitro and in vivo. In addition to inhibiting ribosome biogenesis, CX-5461 causes topoisomerase II trapping and replication-dependent DNA damage, leading to G2/M cell-cycle arrest and apoptotic cell death. Combining CX-5461 with PI does not further enhance the anti-myeloma activity of CX-5461 in vivo. In contrast, CX-5461 shows synergistic interaction with the histone deacetylase inhibitor panobinostat in both the Vk∗MYC and the 5T33-KaLwRij mouse models of MM by targeting ribosome biogenesis and protein synthesis through distinct mechanisms. Our findings thus provide strong evidence to facilitate the clinical development of targeting the ribosome to treat relapsed and refractory MM.
Keywords: CX-5461; MT: Regular Issue; RNA polymerase I; multiple myeloma; panobinostat; ribosome biogenesis.
© 2024 The Author(s).
Conflict of interest statement
R.D.H. is a Chief Scientific Advisor to Pimera, Inc.
Figures



References
-
- Rajkumar S.V., Dimopoulos M.A., Palumbo A., Blade J., Merlini G., Mateos M.V., Kumar S., Hillengass J., Kastritis E., Richardson P., et al. International Myeloma Working Group updated criteria for the diagnosis of multiple myeloma. Lancet Oncol. 2014;15:e538–e548. doi: 10.1016/s1470-2045(14)70442-5. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
