Identifying acute illness phenotypes via deep temporal interpolation and clustering network on physiologic signatures
- PMID: 38600110
- PMCID: PMC11006654
- DOI: 10.1038/s41598-024-59047-x
Identifying acute illness phenotypes via deep temporal interpolation and clustering network on physiologic signatures
Abstract
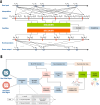
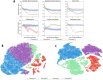
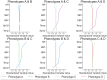
Using clustering analysis for early vital signs, unique patient phenotypes with distinct pathophysiological signatures and clinical outcomes may be revealed and support early clinical decision-making. Phenotyping using early vital signs has proven challenging, as vital signs are typically sampled sporadically. We proposed a novel, deep temporal interpolation and clustering network to simultaneously extract latent representations from irregularly sampled vital signs and derive phenotypes. Four distinct clusters were identified. Phenotype A (18%) had the greatest prevalence of comorbid disease with increased prevalence of prolonged respiratory insufficiency, acute kidney injury, sepsis, and long-term (3-year) mortality. Phenotypes B (33%) and C (31%) had a diffuse pattern of mild organ dysfunction. Phenotype B's favorable short-term clinical outcomes were tempered by the second highest rate of long-term mortality. Phenotype C had favorable clinical outcomes. Phenotype D (17%) exhibited early and persistent hypotension, high incidence of early surgery, and substantial biomarker incidence of inflammation. Despite early and severe illness, phenotype D had the second lowest long-term mortality. After comparing the sequential organ failure assessment scores, the clustering results did not simply provide a recapitulation of previous acuity assessments. This tool may impact triage decisions and have significant implications for clinical decision-support under time constraints and uncertainty.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

References
-
- Hall, M. J., Levant, S. & DeFrances, C. J. Trends in inpatient hospital deaths: National Hospital Discharge Survey, 2000–2010. NCHS Data Brief, 1–8 (2013). - PubMed
-
- Weiss, A. J. & Elixhauser, A. Overview of hospital stays in the United States, 2012. In HCUP Statistical Brief #180 (Agency for Healthcare Research and Quality, 2014). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

