Contribution of a C-Terminal Extension to the Substrate Affinity and Oligomeric Stability of Aldehyde Dehydrogenase from Thermus thermophilus HB27
- PMID: 38602394
- PMCID: PMC11080044
- DOI: 10.1021/acs.biochem.3c00698
Contribution of a C-Terminal Extension to the Substrate Affinity and Oligomeric Stability of Aldehyde Dehydrogenase from Thermus thermophilus HB27
Abstract
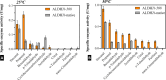
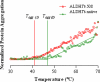
Aldehyde dehydrogenase enzymes (ALDHs) are widely studied for their roles in disease propagation and cell metabolism. Their use in biocatalysis applications, for the conversion of aldehydes to carboxylic acids, has also been recognized. Understanding the structural features and functions of both prokaryotic and eukaryotic ALDHs is key to uncovering novel applications of the enzyme and probing its role in disease propagation. The thermostable enzyme ALDHTt originating fromThermus thermophilus, strain HB27, possesses a unique extension of its C-terminus, which has been evolutionarily excluded from mesophilic counterparts and other thermophilic enzymes in the same genus. In this work, the thermophilic adaptation is studied by the expression and optimized purification of mutant ALDHTt-508, with a 22-amino acid truncation of the C-terminus. The mutant shows increased activity throughout production compared to native ALDHTt, indicating an opening of the active site upon C-terminus truncation and giving rationale into the evolutionary exclusion of the C-terminal extension from similar thermophilic and mesophilic ALDH proteins. Additionally, the C-terminus is shown to play a role in controlling substrate specificity of native ALDH, particularly in excluding catalysis of certain large and certain aromatic ortho-substituted aldehydes, as well as modulating the protein's pH tolerance by increasing surface charge. Dynamic light scattering and size-exclusion HPLC methods are used to show the role of the C-terminus in ALDHTt oligomeric stability at the cost of catalytic efficiency. Studying the aggregation rate of ALDHTt with and without a C-terminal extension leads to the conclusion that ALDHTt follows a monomolecular reaction aggregation mechanism.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
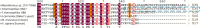


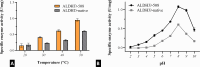

References
-
- Walsh G.Proteins: Biochemistry and Biotechnology; John Wiley & Sons, Incorporated: Hoboken, United Kingdom, 2014, 9781118851494.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

