DArTseq-based SNP markers reveal high genetic diversity among early generation fall armyworm tolerant maize inbred lines
- PMID: 38630672
- PMCID: PMC11023204
- DOI: 10.1371/journal.pone.0294863
DArTseq-based SNP markers reveal high genetic diversity among early generation fall armyworm tolerant maize inbred lines
Abstract
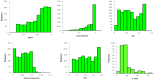
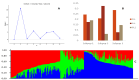
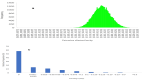
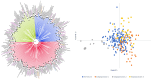
Diversity analysis using molecular markers serves as a powerful tool in unravelling the intricacies of inclusivity within various populations and is an initial step in the assessment of populations and the development of inbred lines for host plant resistance in maize. This study was conducted to assess the genetic diversity and population structure of 242 newly developed S3 inbred lines using 3,305 single nucleotide polymorphism (SNP) markers and to also assess the level of homozygosity achieved in each of the inbred lines. A total of 1,184 SNP markers were found highly informative, with a mean polymorphic information content (PIC) of 0.23. Gene diversity was high among the inbred lines, ranging from 0.04 to 0.50, with an average of 0.27. The residual heterozygosity of the 242 S3 inbred lines averaged 8.8%, indicating moderately low heterozygosity levels among the inbred lines. Eighty-four percent of the 58,322 pairwise kinship coefficients among the inbred lines were near zero (0.00-0.05), with only 0.3% of them above 0.50. These results revealed that many of the inbred lines were distantly related, but none were redundant, suggesting each inbred line had a unique genetic makeup with great potential to provide novel alleles for maize improvement. The admixture-based structure analysis, principal coordinate analysis, and neighbour-joining clustering were concordant in dividing the 242 inbred lines into three subgroups based on the pedigree and selection history of the inbred lines. These findings could guide the effective use of the newly developed inbred lines and their evaluation in quantitative genetics and molecular studies to identify candidate lines for breeding locally adapted fall armyworm tolerant varieties in Ghana and other countries in West and Central Africa.
Copyright: © 2024 Adu et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
Genetic diversity and population structure of early and extra-early maturing maize germplasm adapted to sub-Saharan Africa.BMC Plant Biol. 2021 Feb 17;21(1):96. doi: 10.1186/s12870-021-02829-6. BMC Plant Biol. 2021. PMID: 33596835 Free PMC article.
-
Genetic variation and population structure of maize inbred lines adapted to the mid-altitude sub-humid maize agro-ecology of Ethiopia using single nucleotide polymorphic (SNP) markers.BMC Genomics. 2017 Oct 12;18(1):777. doi: 10.1186/s12864-017-4173-9. BMC Genomics. 2017. PMID: 29025420 Free PMC article.
-
SNP-based assessment of genetic purity and diversity in maize hybrid breeding.PLoS One. 2021 Aug 3;16(8):e0249505. doi: 10.1371/journal.pone.0249505. eCollection 2021. PLoS One. 2021. PMID: 34343170 Free PMC article.
-
Genetic diversity and population structure of maize inbred lines using phenotypic traits and single nucleotide polymorphism (SNP) markers.Sci Rep. 2023 Oct 19;13(1):17851. doi: 10.1038/s41598-023-44961-3. Sci Rep. 2023. PMID: 37857752 Free PMC article.
-
Molecular characterization of diverse CIMMYT maize inbred lines from eastern and southern Africa using single nucleotide polymorphic markers.BMC Genomics. 2012 Mar 25;13:113. doi: 10.1186/1471-2164-13-113. BMC Genomics. 2012. PMID: 22443094 Free PMC article.
Cited by
-
Assessment of fall armyworm tolerant maize hybrids for sustainable maize production in sub-Saharan Africa.Phytoparasitica. 2025;53(2):29. doi: 10.1007/s12600-025-01253-y. Epub 2025 Feb 17. Phytoparasitica. 2025. PMID: 39974150 Free PMC article.
References
-
- Sandhu KS, Singh N. Some properties of corn starches II: Physicochemical, gelatinization, retrogradation, pasting and gel textural properties. Food Chem. 2007;101: 1499–1507. doi: 10.1016/j.foodchem.2006.01.060 - DOI
-
- Shah RT, Prasad K, Kumar P. Maize—A potential source of human nutrition and health: A review. Cogent Food Agric. 2016;1–9. doi: 10.1080/23311932.2016.1166995 - DOI
-
- Cairns JE, Chamberlin J, Rutsaert P, Voss RC, Ndhlela T, Magorokosho C. Challenges for sustainable maize production of smallholder farmers in sub-Saharan Africa. J Cereal Sci. 2021;101: 103274. doi: 10.1016/J.JCS.2021.103274 - DOI
-
- Ekpa O, Palacios-Rojas N, Kruseman G, Fogliano V, Linnemann AR. Sub-Saharan African Maize-Based Foods—Processing Practices, Challenges and Opportunities. 2019;35: 609–639. doi: 101080/875591292019158829010.1080/87559129.2019.1588290
MeSH terms
LinkOut - more resources
Full Text Sources

