Germline homozygosity and allelic imbalance of HLA-I are common in esophagogastric adenocarcinoma and impair the repertoire of immunogenic peptides
- PMID: 38631707
- PMCID: PMC11029431
- DOI: 10.1136/jitc-2023-007268
Germline homozygosity and allelic imbalance of HLA-I are common in esophagogastric adenocarcinoma and impair the repertoire of immunogenic peptides
Abstract
Background: The individual HLA-I genotype is associated with cancer, autoimmune diseases and infections. This study elucidates the role of germline homozygosity or allelic imbalance of HLA-I loci in esophago-gastric adenocarcinoma (EGA) and determines the resulting repertoires of potentially immunogenic peptides.
Methods: HLA genotypes and sequences of either (1) 10 relevant tumor-associated antigens (TAAs) or (2) patient-specific mutation-associated neoantigens (MANAs) were used to predict good-affinity binders using an in silico approach for MHC-binding (www.iedb.org). Imbalanced or lost expression of HLA-I-A/B/C alleles was analyzed by transcriptome sequencing. FluoroSpot assays and TCR sequencing were used to determine peptide-specific T-cell responses.
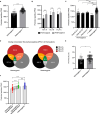
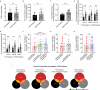
Results: We show that germline homozygosity of HLA-I genes is significantly enriched in EGA patients (n=80) compared with an HLA-matched reference cohort (n=7605). Whereas the overall mutational burden is similar, the repertoire of potentially immunogenic peptides derived from TAAs and MANAs was lower in homozygous patients. Promiscuity of peptides binding to different HLA-I molecules was low for most TAAs and MANAs and in silico modeling of the homozygous to a heterozygous HLA genotype revealed normalized peptide repertoires. Transcriptome sequencing showed imbalanced expression of HLA-I alleles in 75% of heterozygous patients. Out of these, 33% showed complete loss of heterozygosity, whereas 66% had altered expression of only one or two HLA-I molecules. In a FluoroSpot assay, we determined that peptide-specific T-cell responses against NY-ESO-1 are derived from multiple peptides, which often exclusively bind only one HLA-I allele.
Conclusion: The high frequency of germline homozygosity in EGA patients suggests reduced cancer immunosurveillance leading to an increased cancer risk. Therapeutic targeting of allelic imbalance of HLA-I molecules should be considered in EGA.
Keywords: Antigen Presentation; Antigens; Computational Biology; Gastrointestinal Neoplasms; Tumor Escape.
© Author(s) (or their employer(s)) 2024. Re-use permitted under CC BY-NC. No commercial re-use. See rights and permissions. Published by BMJ.
Conflict of interest statement
Competing interests: MvB-B: Honoraria for advisory boards, for invited talks from BMS and financial support for research projects from Astellas, Roche and MSD. HAS: Financial support for research projects from Astra Zeneca. All other authors declare no conflicts of interest.
Figures




Similar articles
-
Immune responses against shared antigens are common in esophago-gastric cancer and can be enhanced using CD40-activated B cells.J Immunother Cancer. 2022 Dec;10(12):e005200. doi: 10.1136/jitc-2022-005200. J Immunother Cancer. 2022. PMID: 36600602 Free PMC article.
-
HLA class I loss in metachronous metastases prevents continuous T cell recognition of mutated neoantigens in a human melanoma model.Oncotarget. 2017 Apr 25;8(17):28312-28327. doi: 10.18632/oncotarget.16048. Oncotarget. 2017. PMID: 28423700 Free PMC article.
-
Identification of a naturally processed NY-ESO-1 peptide recognized by CD8+ T cells in the context of HLA-B51.Cancer Immun. 2002 Sep 19;2:12. Cancer Immun. 2002. PMID: 12747757
-
Neoantigens in Hematological Malignancies-Ultimate Targets for Immunotherapy?Front Immunol. 2019 Dec 20;10:3004. doi: 10.3389/fimmu.2019.03004. eCollection 2019. Front Immunol. 2019. PMID: 31921218 Free PMC article. Review.
-
Immunogenic self-peptides - the great unknowns in autoimmunity: Identifying T-cell epitopes driving the autoimmune response in autoimmune diseases.Front Immunol. 2023 Jan 9;13:1097871. doi: 10.3389/fimmu.2022.1097871. eCollection 2022. Front Immunol. 2023. PMID: 36700227 Free PMC article. Review.
Cited by
-
Neoantigen load as a predictor of relapse in early-stage NSCLC: features that agonise and antagonise prognosis.Cancer Immunol Immunother. 2025 Aug 6;74(9):285. doi: 10.1007/s00262-025-04131-y. Cancer Immunol Immunother. 2025. PMID: 40770517 Free PMC article.
-
Computation strategies and clinical applications in neoantigen discovery towards precision cancer immunotherapy.Biomark Res. 2025 Jul 9;13(1):96. doi: 10.1186/s40364-025-00808-9. Biomark Res. 2025. PMID: 40629481 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
