Preclinical evaluation and first-in-dog clinical trials of PBMC-expanded natural killer cells for adoptive immunotherapy in dogs with cancer
- PMID: 38631708
- PMCID: PMC11029326
- DOI: 10.1136/jitc-2023-007963
Preclinical evaluation and first-in-dog clinical trials of PBMC-expanded natural killer cells for adoptive immunotherapy in dogs with cancer
Abstract
Background: Natural killer (NK) cells are cytotoxic cells capable of recognizing heterogeneous cancer targets without prior sensitization, making them promising prospects for use in cellular immunotherapy. Companion dogs develop spontaneous cancers in the context of an intact immune system, representing a valid cancer immunotherapy model. Previously, CD5 depletion of peripheral blood mononuclear cells (PBMCs) was used in dogs to isolate a CD5dim-expressing NK subset prior to co-culture with an irradiated feeder line, but this can limit the yield of the final NK product. This study aimed to assess NK activation, expansion, and preliminary clinical activity in first-in-dog clinical trials using a novel system with unmanipulated PBMCs to generate our NK cell product.
Methods: Starting populations of CD5-depleted cells and PBMCs from healthy beagle donors were co-cultured for 14 days, phenotype, cytotoxicity, and cytokine secretion were measured, and samples were sequenced using the 3'-Tag-RNA-Seq protocol. Co-cultured human PBMCs and NK-isolated cells were also sequenced for comparative analysis. In addition, two first-in-dog clinical trials were performed in dogs with melanoma and osteosarcoma using autologous and allogeneic NK cells, respectively, to establish safety and proof-of-concept of this manufacturing approach.
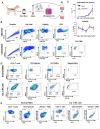
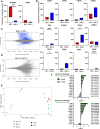
Results: Calculated cell counts, viability, killing, and cytokine secretion were equivalent or higher in expanded NK cells from canine PBMCs versus CD5-depleted cells, and immune phenotyping confirmed a CD3-NKp46+ product from PBMC-expanded cells at day 14. Transcriptomic analysis of expanded cell populations confirmed upregulation of NK activation genes and related pathways, and human NK cells using well-characterized NK markers closely mirrored canine gene expression patterns. Autologous and allogeneic PBMC-derived NK cells were successfully expanded for use in first-in-dog clinical trials, resulting in no serious adverse events and preliminary efficacy data. RNA sequencing of PBMCs from dogs receiving allogeneic NK transfer showed patient-unique gene signatures with NK gene expression trends in response to treatment.
Conclusions: Overall, the use of unmanipulated PBMCs appears safe and potentially effective for canine NK immunotherapy with equivalent to superior results to CD5 depletion in NK expansion, activation, and cytotoxicity. Our preclinical and clinical data support further evaluation of this technique as a novel platform for optimizing NK immunotherapy in dogs.
Keywords: adoptive cell therapy (ACT); clinical trials as topic; computational biology; immunotherapy, adoptive; killer cells, natural.
© Author(s) (or their employer(s)) 2024. Re-use permitted under CC BY-NC. No commercial re-use. See rights and permissions. Published by BMJ.
Conflict of interest statement
Competing interests: None declared.
Figures





Similar articles
-
Radiotherapy enhances natural killer cell cytotoxicity and localization in pre-clinical canine sarcomas and first-in-dog clinical trial.J Immunother Cancer. 2017 Dec 19;5(1):98. doi: 10.1186/s40425-017-0305-7. J Immunother Cancer. 2017. PMID: 29254507 Free PMC article. Clinical Trial.
-
Blood and tissue biomarker analysis in dogs with osteosarcoma treated with palliative radiation and intra-tumoral autologous natural killer cell transfer.PLoS One. 2020 Feb 21;15(2):e0224775. doi: 10.1371/journal.pone.0224775. eCollection 2020. PLoS One. 2020. PMID: 32084139 Free PMC article. Clinical Trial.
-
Natural Killer Cell Expansion with Autologous Feeder Layer and Anti-CD3 Antibody for Immune Cell Therapy of Hepatocellular Carcinoma.Asian Pac J Cancer Prev. 2019 Dec 1;20(12):3797-3803. doi: 10.31557/APJCP.2019.20.12.3797. Asian Pac J Cancer Prev. 2019. PMID: 31870124 Free PMC article.
-
Selection and expansion of natural killer cells for NK cell-based immunotherapy.Cancer Immunol Immunother. 2016 Apr;65(4):477-84. doi: 10.1007/s00262-016-1792-y. Epub 2016 Jan 25. Cancer Immunol Immunother. 2016. PMID: 26810567 Free PMC article. Review.
-
Cellular therapy: Adoptive immunotherapy with expanded natural killer cells.Immunol Rev. 2019 Jul;290(1):85-99. doi: 10.1111/imr.12793. Immunol Rev. 2019. PMID: 31355489 Review.
Cited by
-
Multi-omics analysis of canine aging markers and evaluation of stem cell intervention.Commun Biol. 2025 Jun 10;8(1):905. doi: 10.1038/s42003-025-08333-z. Commun Biol. 2025. PMID: 40494876 Free PMC article.
-
Human immuno-therapeutics for cancer treatment of dogs?Front Vet Sci. 2025 Jul 2;12:1593333. doi: 10.3389/fvets.2025.1593333. eCollection 2025. Front Vet Sci. 2025. PMID: 40671820 Free PMC article. Review.
-
Comparative oncology in action: vignettes on immunotherapy development.Vet Oncol. 2025;2(1):5. doi: 10.1186/s44356-025-00017-4. Epub 2025 Feb 14. Vet Oncol. 2025. PMID: 39958231 Free PMC article. Review.
-
Single cell atlas of canine natural killer cells identifies distinct circulating and tissue resident gene profiles.Front Immunol. 2025 May 15;16:1571085. doi: 10.3389/fimmu.2025.1571085. eCollection 2025. Front Immunol. 2025. PMID: 40443661 Free PMC article.
-
Trained autologous cytotoxic T-cells derived from PBMCs or splenocytes for immunotherapy of neuroblastoma.Front Immunol. 2025 Jun 9;16:1546441. doi: 10.3389/fimmu.2025.1546441. eCollection 2025. Front Immunol. 2025. PMID: 40552293 Free PMC article.
