SARS-CoV-2 remodels the landscape of small non-coding RNAs with infection time and symptom severity
- PMID: 38632240
- PMCID: PMC11024147
- DOI: 10.1038/s41540-024-00367-z
SARS-CoV-2 remodels the landscape of small non-coding RNAs with infection time and symptom severity
Abstract
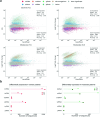
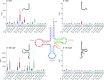
The COVID-19 pandemic caused by the coronavirus SARS-CoV-2 has significantly impacted global health, stressing the necessity of basic understanding of the host response to this viral infection. In this study, we investigated how SARS-CoV-2 remodels the landscape of small non-coding RNAs (sncRNA) from a large collection of nasopharyngeal swab samples taken at various time points from patients with distinct symptom severity. High-throughput RNA sequencing analysis revealed a global alteration of the sncRNA landscape, with abundance peaks related to species of 21-23 and 32-33 nucleotides. Host-derived sncRNAs, including microRNAs (miRNAs), transfer RNA-derived small RNAs (tsRNAs), and small nucleolar RNA-derived small RNAs (sdRNAs) exhibited significant differential expression in infected patients compared to controls. Importantly, miRNA expression was predominantly down-regulated in response to SARS-CoV-2 infection, especially in patients with severe symptoms. Furthermore, we identified specific tsRNAs derived from Glu- and Gly-tRNAs as major altered elements upon infection, with 5' tRNA halves being the most abundant species and suggesting their potential as biomarkers for viral presence and disease severity prediction. Additionally, down-regulation of C/D-box sdRNAs and altered expression of tinyRNAs (tyRNAs) were observed in infected patients. These findings provide valuable insights into the host sncRNA response to SARS-CoV-2 infection and may contribute to the development of further diagnostic and therapeutic strategies in the clinic.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Small non-coding RNA signatures in atrial appendages of patients with atrial fibrillation.J Cell Mol Med. 2024 Jun;28(12):e18483. doi: 10.1111/jcmm.18483. J Cell Mol Med. 2024. PMID: 39051629 Free PMC article.
-
Changes of Small Non-coding RNAs by Severe Acute Respiratory Syndrome Coronavirus 2 Infection.Front Mol Biosci. 2022 Feb 23;9:821137. doi: 10.3389/fmolb.2022.821137. eCollection 2022. Front Mol Biosci. 2022. PMID: 35281271 Free PMC article.
-
In silico approach for the identification of tRNA-derived small non-coding RNAs in SARS-CoV infection.J Appl Genet. 2024 May;65(2):403-413. doi: 10.1007/s13353-024-00853-4. Epub 2024 Mar 21. J Appl Genet. 2024. PMID: 38514586
-
Endogenous miRNA-Based Innate-Immunity against SARS-CoV-2 Invasion of the Brain.Int J Mol Sci. 2023 Feb 8;24(4):3363. doi: 10.3390/ijms24043363. Int J Mol Sci. 2023. PMID: 36834773 Free PMC article. Review.
-
Emerging role of microRNAs and long non-coding RNAs in COVID-19 with implications to therapeutics.Gene. 2023 Apr 20;861:147232. doi: 10.1016/j.gene.2023.147232. Epub 2023 Feb 2. Gene. 2023. PMID: 36736508 Free PMC article. Review.
Cited by
-
Reduced Presence of SARS-CoV-2 microRNA-like Small RNA in the Serum of Patients with Post-Acute Sequelae SARS-CoV-2 Infection.Microorganisms. 2025 Jan 9;13(1):126. doi: 10.3390/microorganisms13010126. Microorganisms. 2025. PMID: 39858894 Free PMC article.
-
LncRNAs Ride the Storm of Epigenetic Marks.Genes (Basel). 2025 Mar 6;16(3):313. doi: 10.3390/genes16030313. Genes (Basel). 2025. PMID: 40149464 Free PMC article. Review.
References
-
- Gorbalenya, A. E. et al. Severe acute respiratory syndrome-related coronavirus: The species and its viruses–a statement of the Coronavirus Study Group. BioRxiv (2020).
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

