T-cell commitment inheritance-an agent-based multi-scale model
- PMID: 38632273
- PMCID: PMC11024127
- DOI: 10.1038/s41540-024-00368-y
T-cell commitment inheritance-an agent-based multi-scale model
Abstract
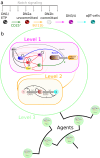
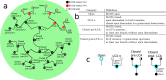
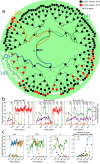
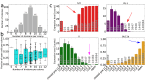
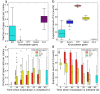
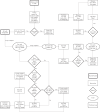
T-cell development provides an excellent model system for studying lineage commitment from a multipotent progenitor. The intrathymic development process has been thoroughly studied. The molecular circuitry controlling it has been dissected and the necessary steps like programmed shut off of progenitor genes and T-cell genes upregulation have been revealed. However, the exact timing between decision-making and commitment stage remains unexplored. To this end, we implemented an agent-based multi-scale model to investigate inheritance in early T-cell development. Treating each cell as an agent provides a powerful tool as it tracks each individual cell of a simulated T-cell colony, enabling the construction of lineage trees. Based on the lineage trees, we introduce the concept of the last common ancestors (LCA) of committed cells and analyse their relations, both at single-cell level and population level. In addition to simulating wild-type development, we also conduct knockdown analysis. Our simulations predicted that the commitment is a three-step process that occurs on average over several cell generations once a cell is first prepared by a transcriptional switch. This is followed by the loss of the Bcl11b-opposing function approximately two to three generations later. This is when our LCA analysis indicates that the decision to commit is taken even though in general another one to two generations elapse before the cell actually becomes committed by transitioning to the DN2b state. Our results showed that there is decision inheritance in the commitment mechanism.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
Update of
-
T-cell commitment inheritance - an agent-based multi-scale model.bioRxiv [Preprint]. 2023 Oct 20:2023.10.18.562905. doi: 10.1101/2023.10.18.562905. bioRxiv. 2023. Update in: NPJ Syst Biol Appl. 2024 Apr 17;10(1):40. doi: 10.1038/s41540-024-00368-y. PMID: 37905091 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

