Hybrid speciation driven by multilocus introgression of ecological traits
- PMID: 38632397
- PMCID: PMC11041799
- DOI: 10.1038/s41586-024-07263-w
Hybrid speciation driven by multilocus introgression of ecological traits
Abstract
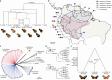
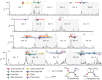
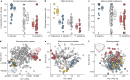
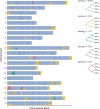
Hybridization allows adaptations to be shared among lineages and may trigger the evolution of new species1,2. However, convincing examples of homoploid hybrid speciation remain rare because it is challenging to demonstrate that hybridization was crucial in generating reproductive isolation3. Here we combine population genomic analysis with quantitative trait locus mapping of species-specific traits to examine a case of hybrid speciation in Heliconius butterflies. We show that Heliconius elevatus is a hybrid species that is sympatric with both parents and has persisted as an independently evolving lineage for at least 180,000 years. This is despite pervasive and ongoing gene flow with one parent, Heliconius pardalinus, which homogenizes 99% of their genomes. The remaining 1% introgressed from the other parent, Heliconius melpomene, and is scattered widely across the H. elevatus genome in islands of divergence from H. pardalinus. These islands contain multiple traits that are under disruptive selection, including colour pattern, wing shape, host plant preference, sex pheromones and mate choice. Collectively, these traits place H. elevatus on its own adaptive peak and permit coexistence with both parents. Our results show that speciation was driven by introgression of ecological traits, and that speciation with gene flow is possible with a multilocus genetic architecture.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






Similar articles
-
Pervasive genetic associations between traits causing reproductive isolation in Heliconius butterflies.Proc Biol Sci. 2011 Feb 22;278(1705):511-8. doi: 10.1098/rspb.2010.1493. Epub 2010 Sep 1. Proc Biol Sci. 2011. PMID: 20810445 Free PMC article.
-
Introgression of wing pattern alleles and speciation via homoploid hybridization in Heliconius butterflies: a review of evidence from the genome.Proc Biol Sci. 2012 Dec 12;280(1752):20122302. doi: 10.1098/rspb.2012.2302. Print 2013 Feb 7. Proc Biol Sci. 2012. PMID: 23235702 Free PMC article. Review.
-
An introgressed wing pattern acts as a mating cue.Evolution. 2015 Jun;69(6):1619-1629. doi: 10.1111/evo.12679. Epub 2015 May 27. Evolution. 2015. PMID: 25930106
-
Review. Hybrid trait speciation and Heliconius butterflies.Philos Trans R Soc Lond B Biol Sci. 2008 Sep 27;363(1506):3047-54. doi: 10.1098/rstb.2008.0065. Philos Trans R Soc Lond B Biol Sci. 2008. PMID: 18579480 Free PMC article. Review.
-
Butterfly genome reveals promiscuous exchange of mimicry adaptations among species.Nature. 2012 Jul 5;487(7405):94-8. doi: 10.1038/nature11041. Nature. 2012. PMID: 22722851 Free PMC article.
Cited by
-
Radiation with reproductive isolation in the near-absence of phylogenetic signal.Sci Adv. 2025 Jul 25;11(30):eadt0973. doi: 10.1126/sciadv.adt0973. Epub 2025 Jul 25. Sci Adv. 2025. PMID: 40712024 Free PMC article.
-
Genetic data suggest gene flow within a narrow hybrid zone between two recently separated species in the genus Parnassius (Lepidoptera: Papilionidae).PLoS One. 2025 Apr 24;20(4):e0321742. doi: 10.1371/journal.pone.0321742. eCollection 2025. PLoS One. 2025. PMID: 40273056 Free PMC article.
-
Evolutionary Histories of Camellia japonica and Camellia rusticana.Ecol Evol. 2024 Dec 24;14(12):e70721. doi: 10.1002/ece3.70721. eCollection 2024 Dec. Ecol Evol. 2024. PMID: 39720636 Free PMC article.
-
Description of a new species of subterranean termite in the genus Reticulitermes (Blattodea: Heterotermitidae) from southern California.Ann Entomol Soc Am. 2025 Jul 3;118(4):315-330. doi: 10.1093/aesa/saaf019. eCollection 2025 Jul. Ann Entomol Soc Am. 2025. PMID: 40708898 Free PMC article.
-
Standing genetic variation and introgression shape the cryptic radiation of Aquilegia in the mountains of Southwest China.Commun Biol. 2025 Apr 30;8(1):684. doi: 10.1038/s42003-025-08120-w. Commun Biol. 2025. PMID: 40307563 Free PMC article.
References
-
- Lamichhaney, S. et al. Rapid hybrid speciation in Darwin’s finches. Science359, 224–228 (2018). - PubMed
-
- Abbott, R. et al. Hybridization and speciation. J. Evol. Biol.26, 229–246 (2013). - PubMed
-
- Schumer, M., Rosenthal, G. G. & Andolfatto, P. How common is homoploid hybrid speciation? Evolution68, 1553–1560 (2014). - PubMed
-
- Lamichhaney, S. et al. Evolution of Darwin’s finches and their beaks revealed by genome sequencing. Nature518, 371–375 (2015). - PubMed
-
- Coyne, J. A. & Orr, H. A. Speciation (Sinauer Associates, 2004).
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

