Endogenous ether lipids differentially promote tumor aggressiveness by regulating the SK3 channel
- PMID: 38642894
- PMCID: PMC11127165
- DOI: 10.1016/j.jlr.2024.100544
Endogenous ether lipids differentially promote tumor aggressiveness by regulating the SK3 channel
Abstract
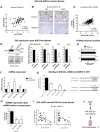
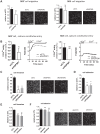
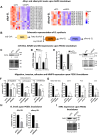
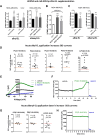
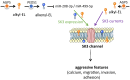
SK3 channels are potassium channels found to promote tumor aggressiveness. We have previously demonstrated that SK3 is regulated by synthetic ether lipids, but the role of endogenous ether lipids is unknown. Here, we have studied the role of endogenous alkyl- and alkenyl-ether lipids on SK3 channels and on the biology of cancer cells. Experiments revealed that the suppression of alkylglycerone phosphate synthase or plasmanylethanolamine desaturase 1, which are key enzymes for alkyl- and alkenyl-ether-lipid synthesis, respectively, decreased SK3 expression by increasing micro RNA (miR)-499 and miR-208 expression, leading to a decrease in SK3-dependent calcium entry, cell migration, and matrix metalloproteinase 9-dependent cell adhesion and invasion. We identified several ether lipids that promoted SK3 expression and found a differential role of alkyl- and alkenyl-ether lipids on SK3 activity. The expressions of alkylglycerone phosphate synthase, SK3, and miR were associated in clinical samples emphasizing the clinical consistency of our observations. To our knowledge, this is the first report showing that ether lipids differentially control tumor aggressiveness by regulating an ion channel. This insight provides new possibilities for therapeutic interventions, offering clinicians an opportunity to manipulate ion channel dysfunction by adjusting the composition of ether lipids.
Keywords: SK3 channel; ether lipids; miRNA; potassium channels.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflict of interests with the contents of article.
Figures

References
-
- Chantome A., Potier-Cartereau M., Clarysse L., Fromont G., Marionneau-Lambot S., Gueguinou M., et al. Pivotal role of the lipid raft SK3-Orai1 complex in human cancer cell migration and bone metastases. Cancer Res. 2013;73:4852–4861. - PubMed
-
- Jaffres P.A., Gajate C., Bouchet A.M., Couthon-Gourves H., Chantome A., Potier-Cartereau M., et al. Alkyl ether lipids, ion channels and lipid raft reorganization in cancer therapy. Pharmacol. Ther. 2016;165:114–131. - PubMed
-
- Diagne A., Fauvel J., Record M., Chap H., Douste-Blazy L. Studies on ether phospholipids. II. Comparative composition of various tissues from human, rat and Guinea pig. Biochim. Biophys. Acta. 1984;793:221–231. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

