Improving the production of carbamoyltobramycin by an industrial Streptoalloteichus tenebrarius through metabolic engineering
- PMID: 38643456
- PMCID: PMC11033246
- DOI: 10.1007/s00253-024-13141-2
Improving the production of carbamoyltobramycin by an industrial Streptoalloteichus tenebrarius through metabolic engineering
Abstract
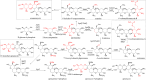
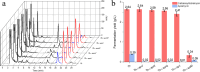
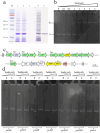
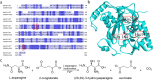
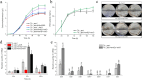
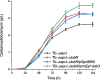
Tobramycin is an essential and extensively used broad-spectrum aminoglycoside antibiotic obtained through alkaline hydrolysis of carbamoyltobramycin, one of the fermentation products of Streptoalloteichus tenebrarius. To simplify the composition of fermentation products from industrial strain, the main byproduct apramycin was blocked by gene disruption and constructed a mutant mainly producing carbamoyltobramycin. The generation of antibiotics is significantly affected by the secondary metabolism of actinomycetes which could be controlled by modifying the pathway-specific regulatory proteins within the cluster. Within the tobramycin biosynthesis cluster, a transcriptional regulatory factor TobR belonging to the Lrp/AsnC family was identified. Based on the sequence and structural characteristics, tobR might encode a pathway-specific transcriptional regulatory factor during biosynthesis. Knockout and overexpression strains of tobR were constructed to investigate its role in carbamoyltobramycin production. Results showed that knockout of TobR increased carbamoyltobramycin biosynthesis by 22.35%, whereas its overexpression decreased carbamoyltobramycin production by 10.23%. In vitro electrophoretic mobility shift assay (EMSA) experiments confirmed that TobR interacts with DNA at the adjacent tobO promoter position. Strains overexpressing tobO with ermEp* promoter exhibited 36.36% increase, and tobO with kasOp* promoter exhibited 22.84% increase in carbamoyltobramycin titer. When the overexpressing of tobO and the knockout of tobR were combined, the production of carbamoyltobramycin was further enhanced. In the shake-flask fermentation, the titer reached 3.76 g/L, which was 42.42% higher than that of starting strain. Understanding the role of Lrp/AsnC family transcription regulators would be useful for other antibiotic biosynthesis in other actinomycetes. KEY POINTS: • The transcriptional regulator TobR belonging to the Lrp/AsnC family was identified. • An oxygenase TobO was identified within the tobramycin biosynthesis cluster. • TobO and TobR have significant effects on the synthesis of carbamoyltobramycin.
Keywords: Apramycin; Biosynthesis; Carbamoyltobramycin; Lrp/AsnC; Oxygenase; Transcriptional regulator.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Barends DM, Brouwers JCAM, Hulshoff A (1987) Fast precolumn derivatization of aminoglycosides with 1-fluoro-2,4-dinitrobenzene and its application to pharmaceutical analysis. J Pharmaceut Biomed 5(6):613–617. 10.1016/0731-7085(87)80073-0 - PubMed
-
- Bibb MJ, White J, Ward JM, Janssen GR (1994) The messenger-rna for the 23s ribosomal-rna methylase encoded by the erme gene of Saccharopolyspora-Erythraea is translated in the absence of a conventional ribosome-binding site. Mol Microbiol 14(3):533–545. 10.1111/j.1365-2958.1994.tb02187.x - PubMed
-
- Brinkman AB, Dahlke I, Tuininga JE, Lammers T, Dumay V, de Heus E, Lebbink JHG, Thomm M, de Vos WM, van der Oost J (2000) An Lrp-like transcriptional regulator from the archaeon Pyrococcus furiosus is negatively autoregulated. J Biol Chem 275(49):38160–38169. 10.1074/jbc.M005916200 - PubMed
-
- Brinkman AB, Ettema TJG, de Vos WM, van der Oost J (2003) The Lrp family of transcriptional regulators. Mol Microbiol 48(2):287–294. 10.1046/j.1365-2958.2003.03442.x - PubMed
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

