This is a preprint.
Nitric oxide inhibits ten-eleven translocation DNA demethylases to regulate 5mC and 5hmC across the genome
- PMID: 38645113
- PMCID: PMC11030528
- DOI: 10.21203/rs.3.rs-4131804/v1
Nitric oxide inhibits ten-eleven translocation DNA demethylases to regulate 5mC and 5hmC across the genome
Update in
-
Nitric oxide inhibits ten-eleven translocation DNA demethylases to regulate 5mC and 5hmC across the genome.Nat Commun. 2025 Feb 18;16(1):1732. doi: 10.1038/s41467-025-56928-1. Nat Commun. 2025. PMID: 39966373 Free PMC article.
Abstract
DNA methylation at cytosine bases of eukaryotic DNA (5-methylcytosine, 5mC) is a heritable epigenetic mark that can regulate gene expression in health and disease. Enzymes that metabolize 5mC have been well-characterized, yet the discovery of endogenously produced signaling molecules that regulate DNA methyl-modifying machinery have not been described. Herein, we report that the free radical signaling molecule nitric oxide (NO) can directly inhibit the Fe(II)/2-OG-dependent DNA demethylases ten-eleven translocation (TET) and human AlkB homolog 2 (ALKBH2). Physiologic NO concentrations reversibly inhibited TET and ALKBH2 demethylase activity by binding to the mononuclear non-heme iron atom which formed a dinitrosyliron complex (DNIC) preventing cosubstrates (2-OG and O2) from binding. In cancer cells treated with exogenous NO, or cells endogenously synthesizing NO, there was a global increase in 5mC and 5-hydroxymethylcytosine (5hmC) in DNA, the substrates for TET, that could not be attributed to increased DNA methyltransferase activity. 5mC was also elevated in NO-producing cell-line-derived mouse xenograft and patient-derived xenograft tumors. Genome-wide DNA methylome analysis of cells chronically treated with NO (10 days) demonstrated enrichment of 5mC and 5hmC at gene-regulatory loci which correlated to changes in the expression of NO-regulated tumor-associated genes. Regulation of DNA methylation is distinctly different from canonical NO signaling and represents a novel epigenetic role for NO.
Conflict of interest statement
Competing interests J.C.C. is the sole inventor on patent application no. 10420838 entitled “Methods for treating cancer using iNOS-inhibitory compositions” held by Houston Methodist Hospital.
Figures

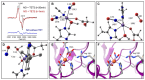



References
-
- Vasudevan D. & Thomas D.D. Insights into the diverse effects of nitric oxide on tumor biology. Vitam Horm 96, 265–98 (2014). - PubMed
-
- Loibl S. et al. The role of early expression of inducible nitric oxide synthase in human breast cancer. Eur J Cancer 41, 265–71 (2005). - PubMed
-
- De Paepe B., Verstraeten V.M., De Potter C.R. & Bullock G.R. Increased angiotensin II type-2 receptor density in hyperplasia, DCIS and invasive carcinoma of the breast is paralleled with increased iNOS expression. Histochem Cell Biol 117, 13–9 (2002). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

