Identifying and Assessing Putative Allosteric Sites and Modulators for CXCR4 Predicted through Network Modeling and Site Identification by Ligand Competitive Saturation
- PMID: 38647430
- PMCID: PMC11139592
- DOI: 10.1021/acs.jpcb.4c00925
Identifying and Assessing Putative Allosteric Sites and Modulators for CXCR4 Predicted through Network Modeling and Site Identification by Ligand Competitive Saturation
Abstract
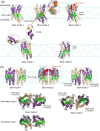
The chemokine receptor CXCR4 is a critical target for the treatment of several cancer types and HIV-1 infections. While orthosteric and allosteric modulators have been developed targeting its extracellular or transmembrane regions, the intramembrane region of CXCR4 may also include allosteric binding sites suitable for the development of allosteric drugs. To investigate this, we apply the Gaussian Network Model (GNM) to the monomeric and dimeric forms of CXCR4 to identify residues essential for its local and global motions located in the hinge regions of the protein. Residue interaction network (RIN) analysis suggests hub residues that participate in allosteric communication throughout the receptor. Mutual residues from the network models reside in regions with a high capacity to alter receptor dynamics upon ligand binding. We then investigate the druggability of these potential allosteric regions using the site identification by ligand competitive saturation (SILCS) approach, revealing two putative allosteric sites on the monomer and three on the homodimer. Two screening campaigns with Glide and SILCS-Monte Carlo docking using FDA-approved drugs suggest 20 putative hit compounds including antifungal drugs, anticancer agents, HIV protease inhibitors, and antimalarial drugs. In vitro assays considering mAB 12G5 and CXCL12 demonstrate both positive and negative allosteric activities of these compounds, supporting our computational approach. However, in vivo functional assays based on the recruitment of β-arrestin to CXCR4 do not show significant agonism and antagonism at a single compound concentration. The present computational pipeline brings a new perspective to computer-aided drug design by combining conformational dynamics based on network analysis and cosolvent analysis based on the SILCS technology to identify putative allosteric binding sites using CXCR4 as a showcase.
Conflict of interest statement
The authors declare the following competing financial interest(s): Alexander D. MacKerell Jr. is co-founder and CSO of SilcsBio LLC.
Figures



Similar articles
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2017 Dec 22;12(12):CD011535. doi: 10.1002/14651858.CD011535.pub2. Cochrane Database Syst Rev. 2017. Update in: Cochrane Database Syst Rev. 2020 Jan 9;1:CD011535. doi: 10.1002/14651858.CD011535.pub3. PMID: 29271481 Free PMC article. Updated.
-
Structure-based discovery of positive allosteric modulators of the A1 adenosine receptor.Proc Natl Acad Sci U S A. 2025 Jul 15;122(28):e2421687122. doi: 10.1073/pnas.2421687122. Epub 2025 Jul 7. Proc Natl Acad Sci U S A. 2025. PMID: 40623180 Free PMC article.
Cited by
-
Allostery Illuminated: Harnessing AI and Machine Learning for Drug Discovery.ACS Med Chem Lett. 2024 Aug 30;15(9):1449-1455. doi: 10.1021/acsmedchemlett.4c00260. eCollection 2024 Sep 12. ACS Med Chem Lett. 2024. PMID: 39291033 Review.
-
Comparative Study of Allosteric GPCR Binding Sites and Their Ligandability Potential.J Chem Inf Model. 2024 Nov 11;64(21):8176-8192. doi: 10.1021/acs.jcim.4c00819. Epub 2024 Oct 23. J Chem Inf Model. 2024. PMID: 39441864 Free PMC article.
-
Exploring Druggable Binding Sites on the Class A GPCRs Using the Residue Interaction Network and Site Identification by Ligand Competitive Saturation.ACS Omega. 2024 Sep 13;9(38):40154-40171. doi: 10.1021/acsomega.4c06172. eCollection 2024 Sep 24. ACS Omega. 2024. PMID: 39346853 Free PMC article.
References
-
- Zhou N.; Luo Z.; Luo J.; Liu D.; Hall J. W.; Pomerantz R. J.; Huang Z. Structural and Functional Characterization of Human CXCR4 as a Chemokine Receptor and HIV-1 Co-Receptor by Mutagenesis and Molecular Modeling Studies. J. Biol. Chem. 2001, 276 (46), 42826–42833. 10.1074/jbc.M106582200. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

